
A Dimensionality Reduction Method for Data Visualization using
Particle Swarm Optimization
Panagiotis C. Petrantonakis
a
and Ioannis Kompatsiaris
b
Information Technologies Institute, Centre for Research and Technology - Hellas (CERTH), Thessaloniki, Greece
Keywords:
Particle Swarm Optimization, Data Visualization, Dimensionality Reduction.
Abstract:
Dimensionality reduction involves mapping of a set of high dimensional input points on a low dimensional
space. Mappings in low dimensional space are expected to preserve the pairwise distances of the high dimen-
sional inputs. In this work we present a dimensionality reduction method, called Dimensionality Reduction
based on Particle Swarm Optimization (PSO-DR), where the conversion of each input to the low dimensional
output does not depend on the rest of the inputs but, instead, it is based on a set of reference points (beacons).
The presented approach results in a simple, fast, versatile dimensionality reduction approach with good quality
of visualization and straightforward out-of-sample extension.
1 INTRODUCTION
In the era of data deluge, robust dimensionality re-
duction (DR) tools for visualization of large, high-
dimensional data have become an imperative need.
The fundamental principle of such tools is to trans-
late high dimensional data so that similar inputs are
mapped onto nearby low dimensional representations.
Ultimately, DR techniques aim at preserving as much
of the high dimensional structure either globally or lo-
cally to the low dimensional representation. A variety
of such tools for DR have been proposed the last few
decades, many of which have been reviewed by Lau-
rens van der Maaten (Van Der Maaten et al., 2009).
DR algorithms fall mainly into two categories.
The ones that preserve the distance-wise global struc-
ture of the data, such as PCA (Hotelling, 1933), Sam-
mon mapping (Sammon, 1969), and others that pre-
serve the structure of a confined neighborhood (local
structure) such as, tSNE (Van Der Maaten and Hin-
ton, 2008), Isomap (Tenenbaum et al., 2000), Lapla-
cian Eigenmaps (Belkin and Niyogi, 2002; Belkin and
Niyogi, 2003), and LLE (Hadsell et al., 2006). Both
categories have been applied in a wide variety of ap-
plications with ever increasing data set sizes. Thus, it
is crucial that DR methods are both versatile and fast,
in order to cope with massive data.
In this paper we introduce a new DR technique for
a
https://orcid.org/0000-0001-9631-4327
b
https://orcid.org/0000-0001-6447-9020
data visualization based on the Particle Swarm Op-
timization algorithm (PSO) (Eberhart and Kennedy,
1995; Shi and Eberhart, 1998; Shi and Eberhart,
1999; Shi et al., 2001). The PSO-based DR ap-
proach (PSO-DR) seeks to preserve the high dimen-
sional structure by exploiting the fast and versatile na-
ture of the PSO algorithm. We provide the respective
algorithmic approach for PSO-DR and present its im-
plementation with parallel, fast computation. We test
PSO-DR on four different datasets, either real or arti-
ficial, and it exhibits good performance on both cate-
gories. We compare the proposed PSO-DR approach
with the current state-of-the-art tSNE method along
with other linear and nonlinear DR algorithms. Its
performance is better or comparable with the most
of the state-of-the-art DR approaches and performs
faster than those with the best visualization quality,
especially for larger data sets. In addition, PSO-DR
allows for the mapping of new data points explic-
itly, in contrast with the majority of the nonlinear DR
techniques where approximate estimation of out-of-
sample extension leads to mapping errors of new data
points. In general, through its simplicity, versatility,
fast computation and straightforward out-of-sample
extension PSO-DR constitutes an efficient, general
purpose, DR technique.
The outline of the paper is as follows. In section
2 we present the methodological approach along with
the background information, PSO-DR algorithm and
implementation issues. Section 3 describes the exper-
imental set up and presents the corresponding results.
Petrantonakis, P. and Kompatsiaris, I.
A Dimensionality Reduction Method for Data Visualization using Particle Swarm Optimization.
DOI: 10.5220/0010020601310138
In Proceedings of the 12th International Joint Conference on Computational Intelligence (IJCCI 2020), pages 131-138
ISBN: 978-989-758-475-6
Copyright
c
2020 by SCITEPRESS – Science and Technology Publications, Lda. All rights reserved
131

Section 4 discusses the presented outcomes and pro-
poses future research. Finally, section 5 concludes the
paper.
2 PSO BASED DATA
VISUALIZATION
2.1 Background
The problem of DR is to find a function that maps
the high dimensional inputs to a lower dimensional
space by preserving the intrinsic structure of the data.
Particularly, we seek to map a high dimensional data
set X ⊂ R
N
to a n-D data set Y ⊂ R
n
in a low di-
mensional space (usually n = 2 or 3, i.e., 2-D or 3-D
space). Each low dimensional point y
i
∈Y, i = 1, ..., M
represents the mapping of a corresponding high di-
mensional point x
i
∈ X , i = 1, ..., M. The approach
followed, e.g., in classical multidimensional scaling
(Torgerson, 1952) is to find all those low dimensional
points y
i
, i = 1, ..., M that minimize the sum of the dif-
ferences between the pairwise distances in the high
dimensional space with the pairwise distances of the
low dimensional one, i.e., to minimize:
φ(Y ) =
∑
i j
(d
2
i j
− δ
2
i j
) (1)
where d
i j
is the distance (dissimilarity measure) be-
tween x
i
and x
j
whereas δ
i j
is the distance of the cor-
responding low dimensional points y
i
and y
j
.
In this work, instead of searching for an optimum
solution set, Y , we seek for the optimum low dimen-
sional points one by one. The key step of the proposed
approach is to define a set of high dimensional bea-
cons, X
b
, i.e., certain, reference points in high dimen-
sional space, and the corresponding low dimensional
ones, Y
b
, i.e., Y
b
is the low dimensional representa-
tion of X
b
. Then, map, e.g., a point x
i
to a point y
i
by
comparing the distances of x
i
from X
b
with the dis-
tances of y
i
from Y
b
. In essence, the relative position-
ing of x
i
with respect to X
b
in high-dimensional space
should be preserved in the low-dimensional space by
regulating the positioning of the y
i
with respect to Y
b
.
The following two subsections describe the PSO al-
gorithm and the proposed PSO-DR approach, respec-
tively.
2.2 Particle Swarm Optimization
Algorithm
PSO algorithm was introduced by Kennedy and Eber-
hart in 1995 (Eberhart and Kennedy, 1995), and it was
inspired by the social behavior of groups of, e.g., birds
in order to solve optimization problems.
PSO algorithm searches the space for optimal so-
lution based on the information shared between the
particles of a group. Each particle follows a trajec-
tory which is influenced by stochastic and determin-
istic components. In particular, each particle moves
according to its best achieved position, in terms of
the optimization problem, and the best position of the
group but with a random component. In every iter-
ation (time point t), a random particle of the group
changes its position, z
t
i
, i = 1, ..., P (P is the popula-
tion of the particles in the group) according to the new
velocity component, v
t
i
(Eberhart and Kennedy, 1995;
Koziel and Yang, 2011), i.e.:
v
t
i
= ωv
t−1
i
+ w
p
r
1
(z
t−1
i, p
− z
t−1
i
) + w
g
r
2
(z
t−1
g
− z
t−1
i
)
(2)
z
t
i
= z
t
i
+ v
t
i
, (3)
where z
t−1
i, p
and z
t−1
g
are the previous best particle
and group positions, respectively, ω, w
p
, w
g
are con-
stant weights and r
1
, r
2
are random numbers. Usually,
search space and velocity values are bounded whereas
the particles are initially distributed randomly in the
search space.
2.3 PSO-DR Algorithm
The proposed approach for DR is based on the PSO
algorithm to find optimal positions Y that correspond
one by one to the high dimensional instances X. Thus,
for the search of an optimal solution y
i
, a group of
particles with positions z
i
, i = 1, ..., P are moving ac-
cording to the rule defined by 2 and 3. The func-
tion that is minimized is the dissimilarity between
the distances of a high dimensional x
i
∈ X from the
high dimensional beacons x
j
b
∈ X
b
, j = 1, ..., J and
the distances of a low dimensional candidate solution
y
i
from the corresponding low dimensional beacons
y
j
b
∈ Y
b
, j = 1, ..., J. In particular we seek to minimize:
φ(y
i
) =
r
∑
j
(d
j
− δ
j
)
2
(4)
where d
j
, j = 1, ..., J are the distances between x
i
and
x
j
b
, j = 1, ..., J, and δ
j
, j = 1, ..., J are the distances be-
tween y
i
and y
j
b
, j = 1, ..., J. In essence, the optimal
solution for Eq. 4, i.e., y
i
, is the mapping of the high
dimensional instance, x
i
, in the low dimensional space
based on the relative distances of x
i
and y
i
from the
corresponding beacon sets. Thus, as soon as the high
and low dimensional beacons are defined, every high
dimensional data can be mapped onto the low dimen-
sional space by minimizing Eq. 4 using PSO.
ECTA 2020 - 12th International Conference on Evolutionary Computation Theory and Applications
132

As high dimensional beacons, X
b
, are used as ref-
erence points, we do not use any particular selection
criterion to define them. Instead, we randomly choose
J instances from data set X to use as the beacon set
X
b
. In order to estimate the corresponding low dimen-
sional beacons, Y
b
, we define as y
1
b
(associated with
the x
1
b
) a zero vector, i.e., y
1
b
= [0...0] ∈ R
n
. Then we
proceed with the definition of the rest of the beacons
y
j
b
, j = 2, ..., J as follows: y
2
b
is estimated by minimiz-
ing φ(y
2
b
) = d
1
− δ
1
where d
1
is the distance between
x
2
b
and x
1
b
, and δ
1
is the distance between the candi-
date y
2
b
and y
1
b
. In accordance, y
3
b
is estimated by min-
imizing φ(y
3
b
) =
q
∑
j
(d
j
− δ
j
)
2
where d
j
, j = 1, 2 are
the respective distances between x
3
b
and x
1
b
, x
2
b
, and
δ
j
, j = 1, 2 are the respective distances between the
candidate y
3
b
and y
1
b
, y
2
b
. The rest of the Y
b
set is esti-
mated with the same procedure. When the whole Y
b
set is defined the rest of the X data set is mapped by
minimizing Eq. 4 with respect to the beacons Y
b
.
2.4 Implementation Issues
The PSO-DR algorithm was implemented in Mat-
lab 2019a. For the PSO algorithm the SwarmOps-
Numerical and Heuristic Optimization toolbox For
Matlab was used (toolbox available at: http://www.
hvass-labs.org/projects/swarmops/matlab/).
We ran all the experiments on a desktop PC, with
Intel Core(TM) i5-9600K at 3.70GHz, and 16 GB of
RAM.
The parameters concerning the PSO algorithm
was chosen according to the best parameters list pre-
sented in (Pedersen, 2010). In particular for the num-
ber of particles (swarm-size, P), number of iterations
of the PSO algorithm (stopping criterion), ω (iner-
tia weight), w
p
(particle’s-best weight), w
g
(swarm’s-
best weight) we used 25, 400, 0.3925, 2.5586, and
1.3358, respectively. These values were used for all
experiments conducted in this work.
For comparison reasons, other DR methods were
also used. In particular, PCA, tSNE, Isomap, Sam-
mon mapping, LLE, and Laplacian Eigenmaps were
compared with PSO-DR. For all these methods the
Matlab Toolbox for Dimensionality Reduction by
Laurens van der Maaten (Van Der Maaten et al., 2009)
was used. For each method the default values of the
parameters provided by the toolbox were used.
The number of beacons for each data set was de-
fined to be a quarter of the number of instances x
i
in each data set X, i.e., J =
1
4
M except for the cases
where J was larger than 1000; then we set J = 1000
irrespectively of the data set size.
3 EXPERIMENTS
The experiments conducted in this work are presented
here. We first describe the data sets used for DR and
subsequently elaborate on the experimental setup. Fi-
nally, we present the respective results.
3.1 Data Sets
Four different data sets were used to eval-
uate the performance of the PSO-DR algo-
rithm. In particular, the MNIST data set (The
MNIST data set is publicly available from
http://yann.lecun.com/exdb/mnist/index.html), the
COIL-20 data set (Nene et al., 1996), the FMNIST
data set (Xiao et al., 2017) and the Swiss Roll data
set with M = 3000.
The MNIST data set contains 60, 000, 28 × 28-
pixel (i.e., N = 784), grayscale images of hand writ-
ten digits (0, ..., 9). In this work we choose randomly
M = 6, 000 images (600 per class) to perform the
experiments. FMNIST data set has the same for-
mat as MNIST except that each class represents fash-
ion items. Again, for FMNIST we choose randomly
M = 6, 000 images (600 per class). The COIL-20 data
set, contains, 32 × 32 (i.e., N = 1, 024) images of 20
different objects which are viewed from 72 orienta-
tions, i.e., resulting in M = 1, 440 images.
3.2 Experimental Setup
For the MNIST, COIL-20, and FMNIST data sets we
use the PSO-DR, PCA, tSNE, Isomap, and Sammon
mapping techniques to transform the high dimen-
sional representations to a two-dimensional (n = 2)
map. For the Swiss Roll data set we use the PSO-DR,
PCA, Laplacian Eigenmaps, Isomap, and LLE tech-
niques to map to the 2-D space. We substituted tSNE
and Sammon mapping with Laplacian Eigenmaps and
LLE as techniques that do not employ neighborhood
graphs perform poorly on the Swiss Roll dataset (Van
Der Maaten et al., 2009).
The resulting maps in each one of the DR task
is shown as a scatter plot. The coloring in the scat-
ter plots is used to provide a way of evaluation for
the performance of the DR techniques. Moreover, for
each one of the DR methods the time needed to map
the respective data set is depicted.
For the proposed PSO-DR method, as soon as the
estimation of the beacons Y
b
is completed, the map-
ping of, e.g., a high dimensional input x
i
to the low di-
mensional space is independent of the mapping of any
other input x
j
, j 6= i. Thus, it is possible to map simul-
taneously multiple inputs. Hence, due to its straight-
A Dimensionality Reduction Method for Data Visualization using Particle Swarm Optimization
133
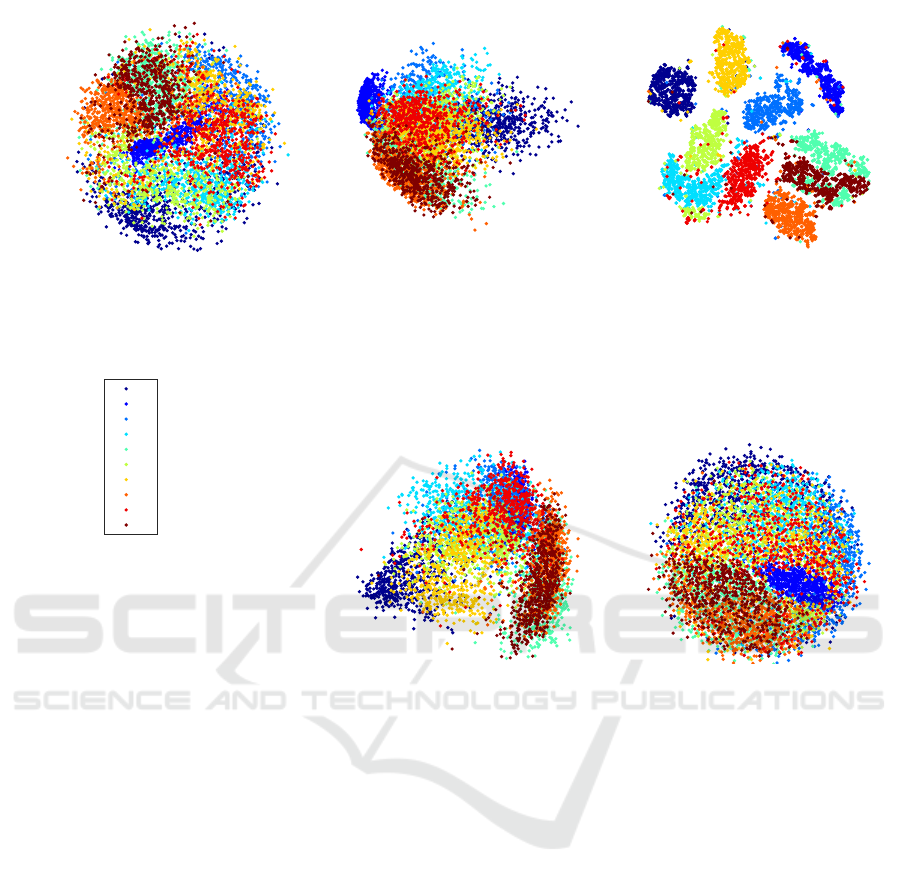
PSO-DR, 57.79 seconds
0
1
2
3
4
5
6
7
8
9
PCA, 1.17 seconds
tSNE, 432.9 seconds
Isomap, 1069.34 seconds
Sammon, 3237.79 seconds
Figure 1: Visualization of the MNIST data set (6, 000 digits) using the PSO-DR, PCA, tSNE, Isomap, and Sammon mapping.
forward parallelization we used all 6 cores of the CPU
to estimate the low dimensional points y
i
in parallel,
accelerating that way the PSO-DR computation.
For the data sets MNIST, COIL-20, and FMINST,
d
i
and δ
i
measures correspond to the Euclidean dis-
tance whereas for Swiss Roll data set δ
i
is the
Euclidean distance whereas d
i
corresponds to the
geodesic distance estimated as in the Isomap method
(Tenenbaum et al., 2000).
3.3 Results
Fig. 1 shows the results of the application of PSO-
DR, PCA, tSNE, Isomap, and Sammon mapping on
the MNIST dataset. PSO-DR algorithm is much faster
than all the rest methods except for PCA. Moreover,
the visualisation quality of the proposed approach
is comparable or better that the majority of the DR
methods apart from tSNE which is currently the best
DR method for data visualization. It is noteworthy
that PSO-DR and Sammon mapping is constructing a
similar ball with PSO-DR exhibiting much faster es-
timation. The mapping of PCA and Isomap exhibit
more extensive overlap between the classes.
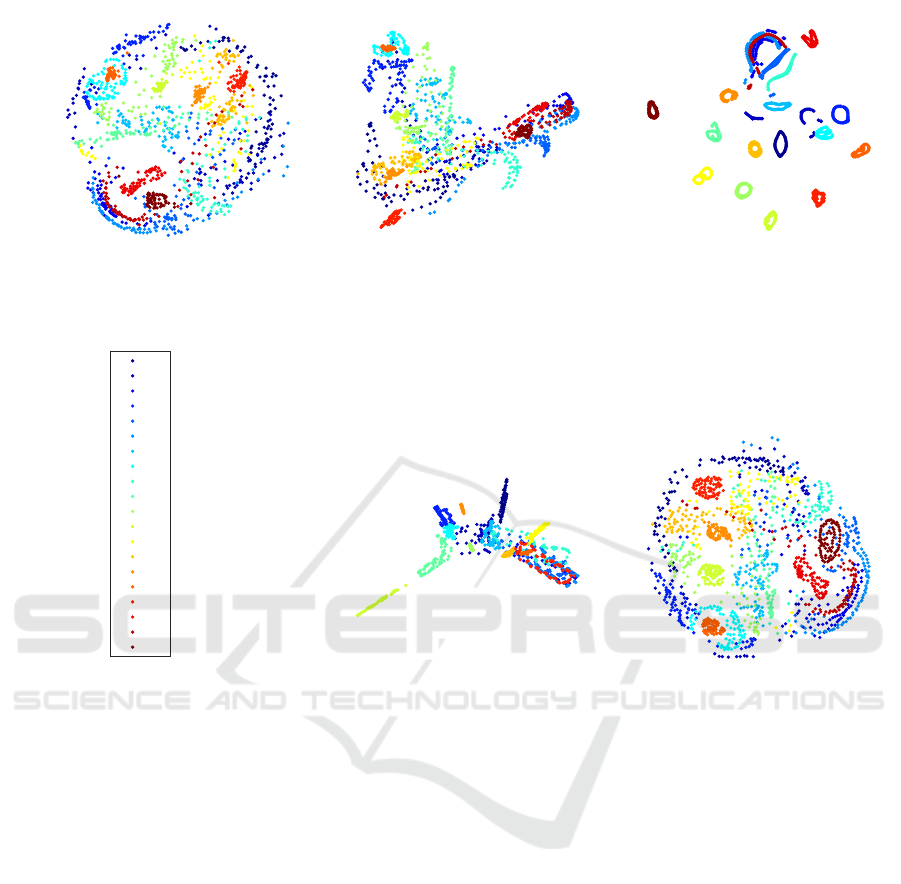
Fig. 2 presents the respective resutls for COIL-20
data set (labels 1-20 refer to each one of the 20 ob-
jects). Again PSO-DR is faster than tSNE and Sam-
mon mapping. Nevertheless, the time differences are
not of the same magnitude and, in addition, PSO-DR
is now slightly slower than Isomap. This is due to
the fact that COIL-20 has much fewer high dimen-
sional points. Thus, PSO-DR’s superiority in terms
of fast computation is mostly revealed in bigger data
sets. Moreover, the similarity of PSO-DR and Sam-
mon mapping is again observed. Nevertheless, Sam-
mon mapping is almost 5 times slower than the PSO-
DR approach.
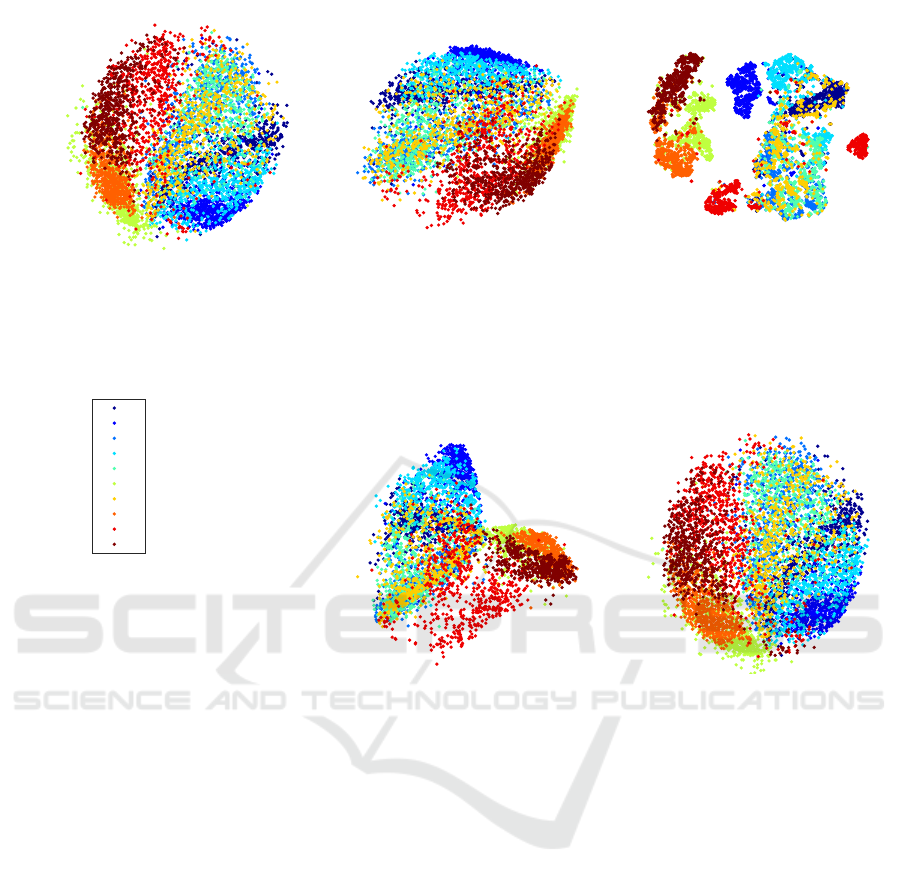
Fig. 3 shows the results of the same DR methods
on FMNIST data set. Similarly, PSO-DR is signifi-
cantly faster from all the other approaches, except for
PCA, with better quality of visualization from PCA
and Isomap, similar quality with Sammon mapping,
and comparable visualization quality with the tSNE
approach.
In Fig. 4 the mappings of the Swiss Roll data
set using the PSO-DR, PCA, Laplacian Eigenmap,
ECTA 2020 - 12th International Conference on Evolutionary Computation Theory and Applications
134

PSO-DR, 18.51 seconds
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
PCA, 0.36 seconds
tSNE, 23.9 seconds
Isomap, 16.77 seconds
Sammon, 90.25 seconds
Figure 2: Visualization of the COIL-20 data set using the PSO-DR, PCA, tSNE, Isomap, and Sammon mapping (labels 1-20
refer to each one of the 20 objects).
Isomap, and LLE methods are presented. In this im-
plementation of the PSO-DR method instead of us-
ing the Euclidean distance as a measure for d
i
, the
geodesic distance as estimated also in Isomap ap-
proach is used. Thus, it is revealed that PSO-DR re-
sults in almost identical representation with Isomap
whereas the rest of the state of the art approaches
have poorer performance. The PSO-DR approach,
though, is slightly slower than Isomap. Nevertheless,
this computation time corresponds to only 6 cores of
parallel computation.
It should be stressed out that if Euclidean distance
is used for the d
i
measure for the case of the Swiss
Roll data set, the performance of the PSO-DR ap-
proach is poorer. Nevertheless, the versatility of the
proposed approach makes it possible to easily adjust
it to the needs of the data set under consideration and
select the dissimilarity measures of the input and out-
put spaces accordingly.
4 DISCUSSION AND FUTURE
WORK
The experiments presented here demonstrate that
PSO-DR is a simple, fast and versatile algorithm for
DR for data visualization where multiple choices for
distance measures both for d
i
and δ
i
are possible in a
simple and straightforward way.
PSO-DR exhibits comparative or better visualiza-
tion quality with the majority of the state of the art
approaches that it was compared to. In essence, apart
from tSNE, PSO-DR outperforms the rest of the ap-
proaches in terms of visualization quality. Neverthe-
less, tSNE is much slower than PSO-DR especially
for large data sets and it performs poorly on certain
datasets like the Swiss Roll dataset. Moreover, for
tSNE, as for many other non-linear DR approaches,
out-of-sample extension is not straightforward (Van
Der Maaten et al., 2009). On the contrary, the out-of-
sample extension in PSO-DR is inherent in its func-
tionality, as any new input can be mapped directly
A Dimensionality Reduction Method for Data Visualization using Particle Swarm Optimization
135

PSO-DR, 54.99 seconds
0
1
2
3
4
5
6
7
8
9
PCA, 0.09 seconds
tSNE, 419.99 seconds
Isomap, 1077.21 seconds
Sammon, 2259.41 seconds
Figure 3: Visualization of the FMNIST data set (6, 000 fashion items) using the PSO-DR, PCA, tSNE, Isomap, and Sammon
mapping. (labeling was used for convenience, Labels 0 to 9 correspond to the 10 different fashion items).
by comparing it with a fixed set of reference beacon-
points.
In addition, the independence of the mapping of
each high dimensional input from other inputs can fa-
vor the straightforward parallel implementation of the
algorithm. Thus, its computation time depends on and
scales in inverse proportion with the number of paral-
lel computation units (cores). Furthermore, the design
of the PSO-DR algorithm makes it a good choice for
ever increasing data sets, even with live streamed data
points as it uses a specified set of beacons to compare
the new data points and map them in the low dimen-
sional space. Nevertheless, in this work, we did not
investigated the functionality of the PSO-DR method
in such cases and we will consider evaluating it in fu-
ture research.
In essence, the only part of the PSO-DR algorithm
where the mapping of an input depends on the previ-
ous inputs is the definition of the beacon set Y
b
. As
soon as this step is completed, PSO-DR mapping can
be performed independently with regard to the differ-
ent inputs. The alternative approach where all map-
pings depend on the previous inputs, thus, no bea-
cons are defined but all x
i
in X are mapped depend-
ing on the distance from all the previous inputs would
result in slower computation without better visualiza-
tion quality. Fig. 5 shows the low dimensional map-
ping of MNIST data set based on such an approach.
The result is comparable with the one shown in Fig. 1
whereas the time needed is multiple times larger that
the one needed with the beacons approach.
5 CONCLUSIONS
In this paper a new approach for DR for data vi-
sualization is presented. The PSO-DR algorithm is
based on the PSO optimization for mapping high di-
mensional inputs to low dimensional spaces with fast
and versatile way with good data visualization qual-
ity. Despite the promising performance of the pro-
posed approach future research will focus on testing
ECTA 2020 - 12th International Conference on Evolutionary Computation Theory and Applications
136
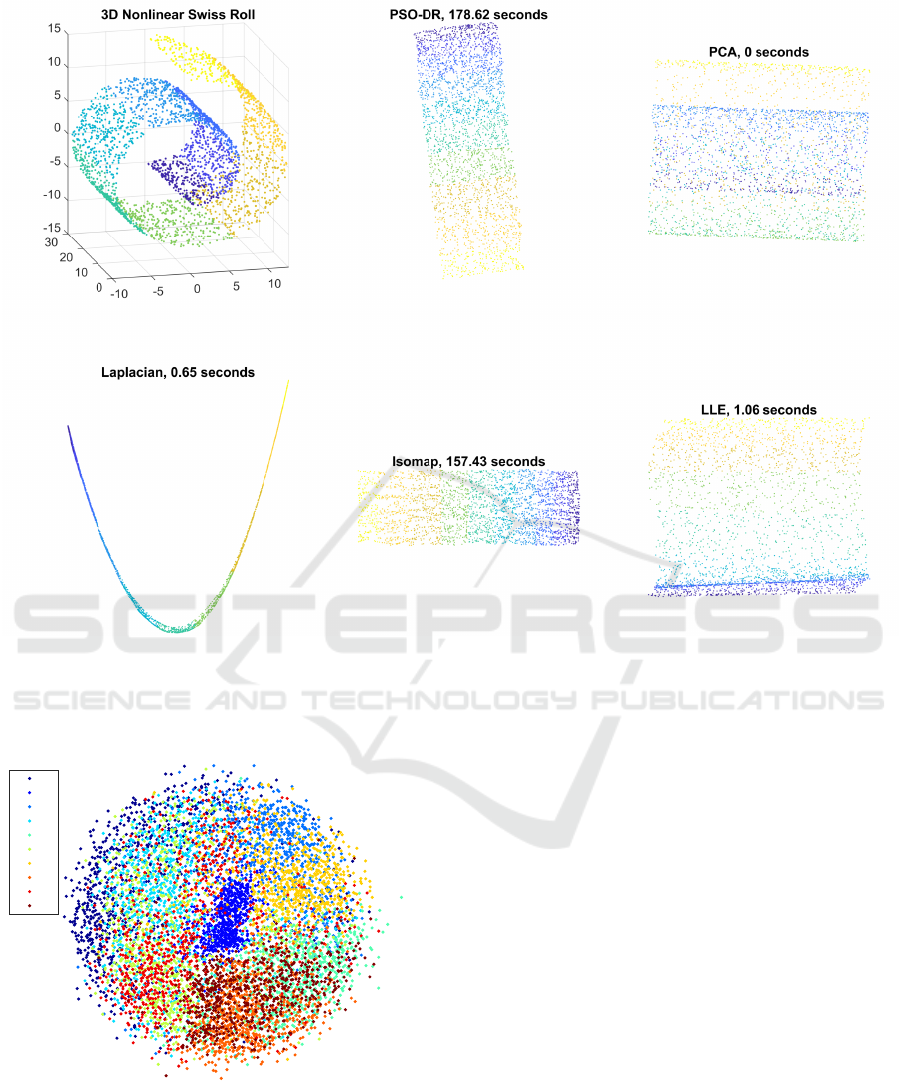
Figure 4: Visualization of the nonlinear Swiss Roll data set in 3D space and by using the PSO-DR, PCA, Laplacian Eigenmaps,
Isomap, and LLE.
PSO-DR, 187.47 seconds
0
1
2
3
4
5
6
7
8
9
Figure 5: Visualization of the MNIST data set using the
PSO-DR without beacon-points.
with more data sets and especially with live streamed
input points.
ACKNOWLEDGEMENTS
This work was supported by the European Union’s
Horizon 2020 Research and Innovation Program
through the project SUITCEYES under Grant
780814.
REFERENCES
Belkin, M. and Niyogi, P. (2002). Laplacian eigenmaps and
spectral techniques for embedding and clustering. In
Advances in neural information processing systems,
pages 585–591.
Belkin, M. and Niyogi, P. (2003). Laplacian eigenmaps
for dimensionality reduction and data representation.
Neural computation, 15(6):1373–1396.
Eberhart, R. and Kennedy, J. (1995). Particle swarm
optimization. In Proceedings of the IEEE inter-
national conference on neural networks, volume 4,
pages 1942–1948. Citeseer.
Hadsell, R., Chopra, S., and LeCun, Y. (2006). Dimen-
sionality reduction by learning an invariant mapping.
A Dimensionality Reduction Method for Data Visualization using Particle Swarm Optimization
137

In 2006 IEEE Computer Society Conference on Com-
puter Vision and Pattern Recognition (CVPR’06), vol-
ume 2, pages 1735–1742. IEEE.
Hotelling, H. (1933). Analysis of a complex of statistical
variables into principal components. Journal of edu-
cational psychology, 24(6):417.
Koziel, S. and Yang, X.-S. (2011). Computational optimiza-
tion, methods and algorithms, volume 356. Springer.
Nene, S. A., Nayar, S. K., Murase, H., et al. (1996).
Columbia object image library (coil-20).
Pedersen, M. E. H. (2010). Good parameters for particle
swarm optimization. Hvass Lab., Copenhagen, Den-
mark, Tech. Rep. HL1001, pages 1551–3203.
Sammon, J. W. (1969). A nonlinear mapping for data
structure analysis. IEEE Transactions on computers,
100(5):401–409.
Shi, Y. and Eberhart, R. C. (1998). Parameter selection in
particle swarm optimization. In International confer-
ence on evolutionary programming, pages 591–600.
Springer.
Shi, Y. and Eberhart, R. C. (1999). Empirical study of par-
ticle swarm optimization. In Proceedings of the 1999
Congress on Evolutionary Computation-CEC99 (Cat.
No. 99TH8406), volume 3, pages 1945–1950. IEEE.
Shi, Y. et al. (2001). Particle swarm optimization: develop-
ments, applications and resources. In Proceedings of
the 2001 congress on evolutionary computation (IEEE
Cat. No. 01TH8546), volume 1, pages 81–86. IEEE.
Tenenbaum, J. B., De Silva, V., and Langford, J. C. (2000).
A global geometric framework for nonlinear dimen-
sionality reduction. Science, 290(5500):2319–2323.
Torgerson, W. S. (1952). Multidimensional scaling: I. the-
ory and method. Psychometrika, 17(4):401–419.
Van Der Maaten, L. and Hinton, G. (2008). Visualizing data
using t-sne. Journal of machine learning research,
9(Nov):2579–2605.
Van Der Maaten, L., Postma, E., and Van den Herik, J.
(2009). Dimensionality reduction: a comparative re-
view. J Mach Learn Res, 10(66-71):13.
Xiao, H., Rasul, K., and Vollgraf, R. (2017). Fashion-
mnist: a novel image dataset for benchmark-
ing machine learning algorithms. arXiv preprint
arXiv:1708.07747.
ECTA 2020 - 12th International Conference on Evolutionary Computation Theory and Applications
138
