
Non-linear Transformations of Vector Space Embedded
Graphs
Kaspar Riesen and Horst Bunke
Institute of Computer Science and Applied Mathematics, University of Bern
Neubr¨uckstrasse 10, CH-3012 Bern, Switzerland
Abstract. In pattern recognition and related areas an emerging trend of repre-
senting objects by graphs can be observed. As a matter of fact, graph based rep-
resentation offers a powerful and flexible alternative to the widely used feature
vectors. However, the space of graphs contains almost no mathematical structure,
and consequently, there is a lack of suitable algorithms for graph classification,
clustering, and analysis. Recently, a general approach to transforming graphs into
n-dimensional real vectors has been proposed. In the present paper we use this
method, which can also be regarded as a novel graph kernel, and investigate the
application of kernel principal component analysis (kPCA) on the resulting vec-
tor space embedded graphs. To this end we consider the common task of object
classification, and show that kPCA in conjunction with the novel graph kernel
outperforms different reference systems on several graph data sets of diverse na-
ture.
1 Introduction
The first step in any system in pattern recognition, machine learning, data mining, and
related fields consists of the representation of objects by adequate data structures. In
statistical approaches the data structure is given by n-dimensional vectors x ∈ R
n
,
where each of the n dimensions represents the value of a specific feature. In recent
years a huge amount of algorithms for classification, clustering, and analysis of objects
given in terms of feature vectors have been developed [1–3].
Yet, the use of feature vectors implicates two limitations. First, as vectors describe a
predefined set of features, all vectors of a set have to preserve the same length regardless
of the size or complexity of the corresponding objects. Furthermore, there is no direct
possibility to describe binary relationships among different parts of an object. It is well
known that both constraints can be overcome by graph based representation [4]. That
is, graphs allow us to adapt their size to the complexity of the underlying object and
they offer a convenient possibility to describe relationships among different parts of an
object.
The major drawback of graphs, however, is that they offer little mathematical struc-
ture, i.e. most of the basic mathematical operations are not available or not defined in
a standardized way for graphs. Nevertheless, since the concept of kernel machines has
been extended from vectors to symbolic data structure, and in particular to graphs [5],
this drawback can be overcome. The key idea of such kernel machines is that rather than
Riesen K. and Bunke H. (2008).
Non-linear Transformations of Vector Space Embedded Graphs.
In Proceedings of the 8th International Workshop on Pattern Recognition in Information Systems, pages 173-183
Copyright
c
SciTePress

defining handcrafted mathematical operations in the original graph domain, the graphs
are mapped into a vector space where all those operations are readily available. Obvi-
ously, by means of graph kernels one can benefit from both the high representational
power and flexibility of graphs and the wide range of pattern recognition algorithms for
feature vectors.
In the present paper we use the dissimilarity space embedding graph kernel [6]. In
order to map individual graphs into a vector space, this graph kernel employs graph edit
distance. Consequently, it can be applied to any kind of graphs (directed or undirected,
labeled or unlabeled). If we allow labels on the nodes or edges, these labels can be
symbolic, numerical or whole attribute vectors. The basic idea of this approach is to
transform a given graph g into a n
′
-dimensional
1
vector x = (x
1
, . . . , x
n
′
), where each
component x
i
(1 ≤ i ≤ n
′
) is equal to the graph edit distance of g to the i-th graph of
a predefined set of prototype graphs P.
In a first attempt to define an appropriate set P, prototype selection methods have
been applied to a training set [6]. Next, a more general approach without heuristic pro-
totype selection methods has been proposed [7]. In this approach the whole training set
T is used as prototype set P, i.e. P = T . Two classical approaches of linear transfor-
mations, i.e. Principal Component Analysis (PCA) and Multiple Discriminant Analysis
(MDA) [1], have been applied to vector space embedded graphs subsequently.
In the present paper we build upon this idea, but in the extended fashion of non-
linear transformation by means of Kernel PCA. That is, we transform graphs into vec-
tors by edit distance computation utilizing a whole training set T and eventually apply
kernel PCA to the resulting vectorial descriptions of the graphs. With several experi-
mental results we show that this approach outperforms both a reference system applied
in the original graph domain and a reference system in the embedded vector space in
conjunction with linear PCA.
2 Dissimilarity Space Embedding of Graphs
Similarly to the graph kernels described in [8] the embedding procedure proposed in
this paper makes use of graph edit distance. The key idea of graph edit distance is to
define the dissimilarity, or distance, of graphs by the minimum amount of distortion that
is needed to transform one graph into another. A standard set of distortion operations is
given by insertions, deletions, and substitutions of nodes and edges.
Given two graphs, the source graph g
1
and the target graph g
2
, the idea of graph edit
distance is to delete some nodes and edges from g
1
, relabel (substitute) some of the re-
maining nodes and edges, and insert some nodes and edges in g
2
, such that g
1
is finally
transformed into g
2
. A sequence of edit operations e
1
, . . . , e
k
that transform g
1
into g
2
is called an edit path between g
1
and g
2
. Obviously, for every pair of graphs (g
1
, g
2
),
there exist a number of differentedit paths transforming g
1
into g
2
. Let Υ (g
1
, g
2
) denote
the set of all such edit paths. To find the most suitable edit path out of Υ (g
1
, g
2
), one
introduces a cost for each edit operation, measuring the strength of the corresponding
operation. The idea of such cost functions is to define whether or not an edit operation
1
We use n
′
instead of n for the sake of consistency with the remainder of this paper.

represents a strong modification of the graph. Hence, between two similar graphs, there
should exist an inexpensive edit path, representing low cost operations, while for dif-
ferent graphs an edit path with high costs is needed. Consequently, the edit distance of
two graphs is defined by the minimum cost edit path between two graphs.
Definition 1 (Graph Edit Distance) Let g
1
= (V
1
, E
1
, µ
1
, ν
1
) be the source graph
and g
2
= (V
2
, E
2
, µ
2
, ν
2
) be the target graph. The graph edit distance between g
1
and
g
2
is defined by
d(g
1
, g
2
) = min
(e
1
,...,e
k
)∈Υ (g
1
,g
2
)
k
X
i=1
c(e
i
),
where Υ (g
1
, g
2
) denotes the set of edit paths transforming g
1
into g
2
, and c denotes the
edit cost function measuring the strength c(e
i
) of edit operation e
i
.
The edit distance of graphs can be computed,for example, by a tree search algorithm
[9] or by faster, suboptimal methods which have been proposed recently [10].
2.1 Basic Embedding Approach
Different approaches to graph embedding have been proposed in the literature [11–13].
In [11], for instance, an embedding based on algebraic graph theory and spectral matrix
decomposition is proposed. Applying an error-tolerant string matching algorithm to the
eigensystem of graphs to infer distances of graphs is proposed in [12]. These distances
are then used to embed the graphs into a vector space by multidimensional scaling. In
[13] features derived from the eigendecompostion of graphs are studied. In fact, such
feature extraction defines an embedding of graphs into vector spaces, too.
Recently, it has been proposed to embed graphs in vector spaces by means of
edit distance and prototypes [6]. The idea underlying this method was first developed
for the embedding of real vectors in a dissimilarity space [14]. In this method, after
having selected a set P = {p
1
, . . . , p
n
′
} of n
′
≤ n prototypes from a training set
T = {g
1
, . . . , g
n
}, the dissimilarity of a graph g to each prototype p ∈ P is computed.
This leads to n
′
dissimilarities, d
1
= d(g, p
1
), . . . , d
n
′
= d(g, p
n
′
), which can be in-
terpreted as an n
′
-dimensional vector (d
1
, . . . , d
n
′
). In this way we can transform any
graph from the training set, as well as any other graph from a validation or testing set,
into a vector of real numbers.
Definition 2 (Graph Embedding) Given are a graph space G and a training set of
graphs T = {g
1
, . . . , g
n
} ⊆ G. If P = { p
1
, . . . , p
n
′
} ⊆ T is a set of prototypes, the
mapping t
P
n
′
: G → R
n
′
is defined as a function
t
P
n
′
(g) 7→ (d(g, p
1
), . . . , d(g, p
n
′
))
where d(g, p
i
) is the graph edit distance between the graph g ∈ G and the i-th proto-
type.
One crucial question in this approach is how to find a subset P of prototypes that
lead to a good performance of the classifier in the feature space. As a matter of fact,

both the individual prototypes selected from T and their number have a critical impact
on the classifier’s performance. In [6,14] different prototype selection algorithms are
discussed. It turns out that none of them is globally best, i.e. the quality of a prototype
selector depends on the underlying data set.
In a recent paper it is proposed to use all available elements from the training set
of prototypes, i.e. P = T and subsequently apply dimensionality reduction methods.
This process is much more principled than the previous approaches and completely
avoids the difficult problem of heuristic prototype selection. For dimensionality reduc-
tion Principal Component Analysis (PCA) and Fisher’s Linear Discriminant Analysis
(LDA) [1, 15] are used. In the present paper we use the same idea but with an exten-
sion of PCA to non-linear distributions. That is, instead of directly applying a PCA to
the vectors resulting from mapping t
P
n
′
with n
′
= n, the vectors are implicitly mapped
into a higher-dimensional feature space by means of a kernel function. In this higher-
dimensional feature space PCA is then applied to the vector maps. This procedure is
commonly referred to as kernel PCA [16].
3 PCA and Kernel PCA
In this section we first review linear transformations by means of PCA and then describe
a non-linear extension by means of kernel PCA.
3.1 PCA
Principal Component Analysis (PCA) [3] is a linear transformation which basically
seeks the projection that best represents the data. PCA finds a new space whose basis
vectors correspond to the maximum variance directions in the original space. PCA falls
in the category of unsupervised transformation methods, i.e. PCA does not take any
class label information into consideration. Let us assume that m objects are given in
terms of n-dimensional column vectors x ∈ R
n
. We first normalize the data by shifting
the sample mean µ =
1
m
P
m
i=1
x
i
to the origin of the coordinate system, i.e. we center
the data. Next we compute the covariance matrix C of the centered data which is defined
as
C =
1
m
m
X
i=1
x
i
x
′
i
The covariance matrix C is symmetric and, therefore, an orthogonal basis can be
defined by finding the eigenvalues λ
i
and the corresponding eigenvectors e
i
of mC. To
this end the following eigenstructure decomposition has to be solved: mCe
i
= λ
i
e
i
.
The eigenvectors W = (e
1
, . . . , e
n
) are also called principal components and they
build an orthogonal basis. Consequently, the matrix W represents a linear transforma-
tion that maps the original data points x ∈ R
n
to new data points y ∈ R
n
where
y = W
′
x
That is, the data is projected into the space spanned by the eigenvectors. The eigenval-
ues λ
i
represent the variance of the data points in the direction of the corresponding

eigenvector e
i
. Hence, the eigenvectors e
i
can be ordered according to decreasing mag-
nitude of their corresponding eigenvalues. Consequently, the first principal component
points in the direction of the highest variance and, therefore, includes the most infor-
mation about the data. The second principal component is perpendicular to the first
principal component and points in the direction of the second highest variance and so
on. In order to project the n-dimensional data x ∈ R
n
into a subspace of lower dimen-
sionality, we retain only the n
′
< n eigenvectors W
n
′
= (e
1
, . . . , e
n
′
) with the highest
n
′
eigenvalues. Formally, the mapping of x ∈ R
n
to ˜y ∈ R
n
′
is defined by
˜y = W
′
n
′
x
Note that the larger the resulting dimensionality n
′
is defined, the greater is the fraction
of the captured variance.
3.2 Kernel PCA
Kernel machines constitutes a very powerful class of algorithms in the field of pattern
recognition. The idea of kernel machines is to map a vector by means of a kernel func-
tion into a higher-dimensional vector space. This procedure offers a very elegant way
to construct non-linear extensions of linear algorithms in pattern recognition.
Definition 3 (Kernel Function) Let x, y ∈ R
n
be feature vectors and ψ : R
n
→ F
a (possibly nonlinear) function where n ∈ N and F is a (possibly infinite dimen-
sional) feature space. A kernel function is a mapping κ : R
n
× R
n
→ R such that
κ(x, y) = hψ(x), ψ(y)i.
According to the definition above, the result of a kernel function k(x, y), applied to
two feature vectors x and y, is equal to the result that one obtains by mapping those vec-
tors to a possibly infinite dimensional feature space F and computing their dot product
subsequently. The fundamental observation that makes kernel theory so interesting in
the field of pattern recognition is that many of the clustering and classification algo-
rithms can be kernelized, i.e. they can be formulated entirely in terms of dot products.
Consequently, instead of mapping patterns from the original pattern domain to a feature
space and computing the dot product there, one can simply evaluate the value of the
kernel function κ in the original pattern space. This procedure is commonly referred to
as the kernel trick [17,18] because of its property of determining the dot product in a
higher-dimensional feature space immediately without performing the transformation
explicitly.
As a matter of fact, PCA can be reformulated in terms of dot products only, i.e. PCA
is kernelizable [16]. In order to see this, let us first assume that we apply the mapping
ψ : R
n
→ F explicitly to our data. For further processing we assume that the data
is centered in the kernel feature space
2
. We compute the covariance matrix C in the
feature space F
C =
1
m
m
X
i=1
ψ(x
i
)ψ(x
i
)
′
2
We will come back to this point later in this section.

Similarly to PCA the principal components of this feature space are extracted by means
of the eigenstructure decomposition
λw = mCw (1)
Since
mCw =
m
X
i=1
(ψ(x
i
)
′
w)ψ(x
i
)
all solutions w must lie in the span of ψ(x
1
), . . . , ψ(x
m
). This observation is crucial
since it allows us to rewrite w as a linear combination of the vectors ψ(x
i
) with coeffi-
cients α
i
(i = 1, . . . , m)
w =
m
X
i=1
α
i
ψ(x
i
) (2)
Furthermore Equation 1 can be rewritten as
λ
m
X
i=1
α
i
ψ(x
i
) =
m
X
i,j=1
α
i
ψ(x
j
)(ψ(x
j
)
′
ψ(x
i
))
Obviously this is equivalent to m equations (k = 1, . . . , m)
λ
m
X
i=1
α
i
(ψ(x
i
)
′
ψ(x
k
)) =
m
X
i,j=1
α
i
(ψ(x
j
)
′
ψ(x
k
))(ψ(x
j
)
′
ψ(x
i
)) (3)
The tremendous benefit of Equation 3 is that it is entirely formulated in terms of dot
products. Hence, we can define a kernel matrix K by K
i,j
= ψ(x
i
)
′
ψ(x
j
) and replace
the dot products in Equation 3 by the kernel function. Formally, λKα = K
2
α where
α = (α
1
, . . . , α
m
)
′
. K is by definition symmetric and has a set of eigenvectors that
span the whole space. Consequently, all solutions α can be obtained by the eigende-
composition of K: λα = Kα.
Let λ
1
≤ λ
2
≤ . . . ≤ λ
m
denote the eigenvalues and α
1
, . . . α
m
the corresponding
eigenvectors of K. The principal components, i.e. eigenvectors w in the feature space
F, need to be normalized to have unit length. This can be achieved by means of the
kernel matrix K.
||w||
2
=
m
X
i=1
α
i
ψ(x
i
)
!
′
m
X
j=1
α
j
ψ(x
j
)
= α
′
Kα = λα
′
α
Hence, we normalize the coefficients α by 1 = λα
′
α. In order to compute a projection
of ψ(x) ∈ F onto a subspace spanned by the first n
′
eigenvectors W
n
′
the following
equation is used
˜y
k
= W
′
n
′
ψ(x)
=
m
X
i=1
α
k
i
ψ(x
i
)
′
ψ(x)
!
n
′
k=1
=
m
X
i=1
α
k
i
κ(x
i
, x)
!
n
′
k=1

That is, the mapping can be computed by means of the kernel matrix and the normalized
coefficients α
k
only.
So far we assumed that the data is centered. Usually, this is not fulfilled, of course.
In an explicit fashion one would center the data by
ˆ
ψ(x) = ψ(x) −
1
m
m
X
i=1
ψ(x
i
)
However, it turns out that the centering of the data has not to be done explicitly. That is,
the kernel matrix K can be replaced by
ˆ
K which is defined by
ˆ
K
i.j
=
ˆ
ψ(x)
′
ˆ
ψ(y)
=
1
m
m
X
i=1
ψ(x)
′
ψ(x
i
) −
1
m
m
X
i=1
ψ(y)
′
ψ(x
i
) +
1
m
2
m
X
i,j=1
ψ(x
i
)
′
ψ(x
j
)
=
1
m
m
X
i=1
κ(x, x
i
) −
1
m
m
X
i=1
κ(y, x
i
) +
1
m
2
m
X
i,j=1
κ(x
i
, x
j
)
4 Experimental Results
The experiments in the present paper consider the task of object classification on six dif-
ferent graph datasets. We use three reference systems to compare our algorithm with.
First, a nearest neighbor classifier is applied directly in the domain of graphs. Note that
as of today – up to a few exceptions, e.g. in [19] – this is the only classifier directly
applicable to general graphs. The second reference system is a support vector machine
(SVM) applied to the untransformed embedding vectors. The third reference system is
an SVM with RBF-Kernel applied to PCA reduced vectors. The RBF-kernel SVM used
in this paper has parameters C and γ. C corresponds to the weighting factor for misclas-
sification penalty and γ > 0 is used in our kernel function κ(u, v) = exp(−γ||u−v||
2
).
Note that both the number of dimensions n
′
retained by PCA and the SVM parameters
(C, γ) are optimized on an independent validation set.
Our new approach also makes use of the vector space embedded graphs. However,
we apply an RBF-kernel PCA to the resulting vectors instead of a linear transformation.
Then an SVM with linear kernel κ(u, v) = ||u−v|| is applied to the non-linearly trans-
formed and reduced data for the purpose of classification. Hence, besides the number of
dimensions n
′
retained by kernel PCA only the SVM parameter C has to be optimized.
4.1 Datasets
For our experimental evaluation, six data sets with quite different characteristics are
used. The datasets vary with respect to graph size, edge density, type of labels for the
nodes and edges, and meaning of the underlying objects. Lacking space we give a short
description of the data only. For a more thorough description we refer to [6] where
the same datasets are used. Note that all of the datasets are divided into three disjoint
subsets, i.e. a training set, a validation set, and a test set.

The first dataset used in the experiments consists of graphs representing distorted
letter drawings out of 15 classes (LETTER DATABASE). The next dataset consists of
graphs representing distorted symbols from architecture, electronics, and other techni-
cal fields out of 32 different classes [20] (GREC DATABASE). Next we apply the pro-
posed method to the problem of image classification, i.e. we use graphsrepresenting im-
ages out of the four different classes city, countryside, snowy, and people (IMAGE). The
fourth dataset is given by graphs representing fingerprint images of the NIST-4 database
[21] out of the four classes arch, left, right, and whorl (FINGERPRINT DATABASE). The
fifth graph set is constructed from the AIDS Antiviral Screen Database of Active Com-
pounds [22]. Graphs from this database belong to two classes (active, inactive), and rep-
resent molecules with activity against HIV or not (AIDS DATABASE). The last dataset
consists of graphs representing webpages [23] that origin from 20 different categories
(Business, Health, Politics, . . .) (WEB DATABASE).
4.2 Results and Discussion
In Table 1 the classification accuracies on the test sets of all three reference systems (k-
nearest neighbor classifier in the graph domain, SVM applied to unreduced data, SVM
based on the PCA reduced data) and our novel approach (SVM based on kernel PCA
reduced data) are given.
First of all we observe that the SVM based on the unreduced embedding data, SVM
(all), outperforms the first reference system in five out of six cases. PCA-SVM and
kPCA-SVM outperform the first reference system even on all datasets. All performance
improvementsare statistically significant. Obviously,the dissimilarity space embedding
graph kernel in conjunction with SVM is a powerful and robust methodology for graph
classification.
Comparing the SVM applied on unreduced data with the SVM based on PCA re-
duced data, we observe that the latter outperforms the former on five out of six datasets
(twice with statistical significance). That is, the reduction of the data by means of PCA
leads to a further increase in classification accuracy. The SVM in conjunction with
the kPCA reduced data also improves five out of six results compared to the second
reference system (twice with statistical significance). Note that the percentage of the
dimensions retained by kernel PCA is indicated in brackets.
Comparing the classification results achieved with SVM on PCA and kernel PCA
reduced data, we observe that in four out of six cases the accuracy is improved and only
once it deteriorates by the non-linear transformation. Hence, there is a clear tendency
that the new approach with kernel PCA is favorable.
From some higher level of abstraction the last observation reflects the fact that both
systems (PCA-SVM and kPCA-SVM) are quite similar. In Fig. 1 both approaches are
shown schematically. The first system (solid arrows) transforms the graph domain G
into a vector space R
n
. Next linear transformation (PCA) for mapping the data into
an n
′
-dimensional vector space is used. Finally, an RBF-kernel support vector machine
is applied which implicitly maps the graphs into a (possibly infinite) feature space F
2
and labels the objects according to a label space Ω. The second system (dashed ar-
rows), however, uses non-linear transformation of the data (kPCA via F
1
to R
n
′
) in
conjunction with linear support vector machine. So both systems consist of a linear and
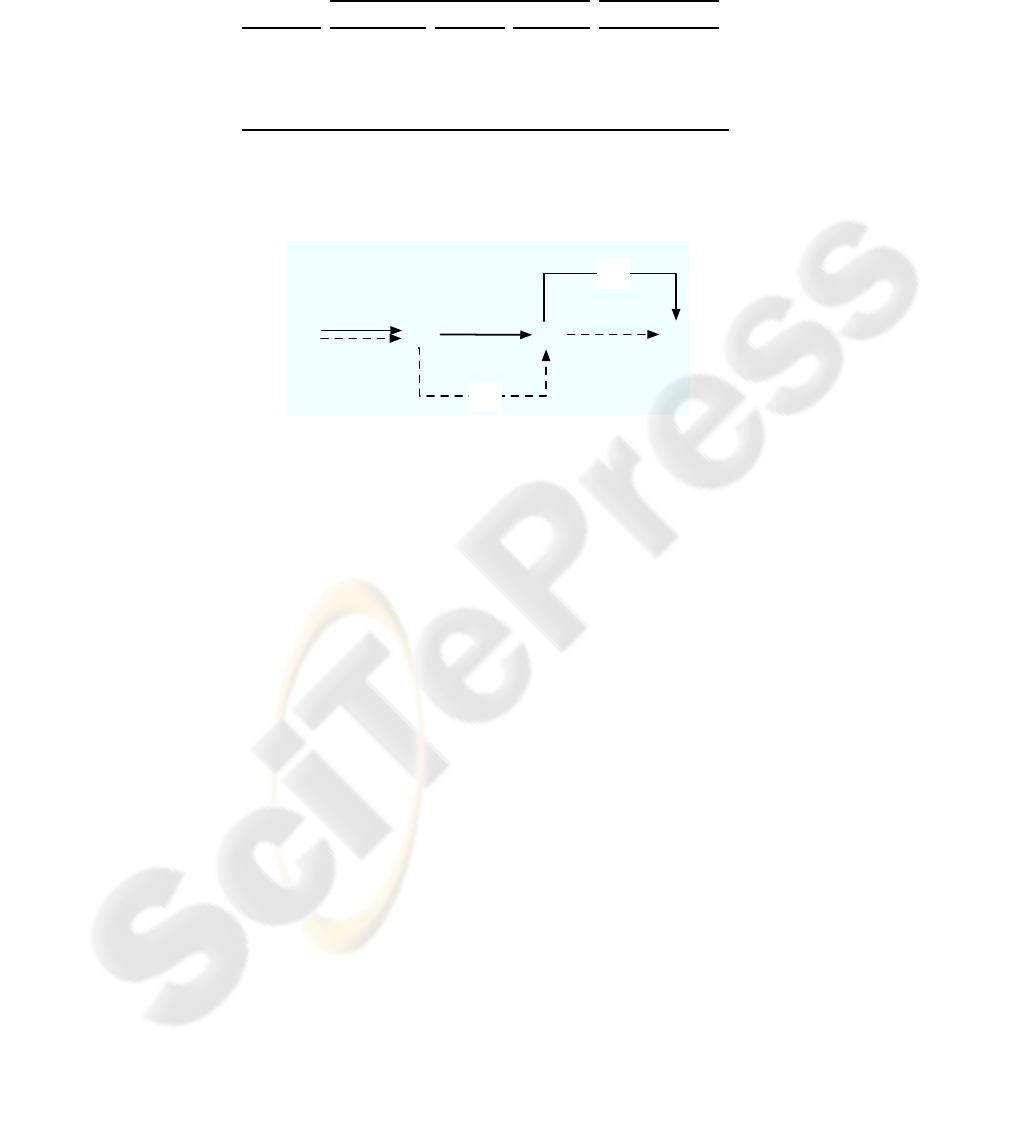
Table 1. Classification accuracy in the original graph domain and in the embedded
vector space.
Reference Systems Proposed System
Database k-NN (graph) SVM (all) PCA-SVM kPCA-SVM
LETTER 91.1 92.3 ① 92.7 ① 93.2 (6%) ①
GREC 86.0 91.3 ① 92.9 ① 95.0 (25%) ①②③
IMAGE 59.5 75.2 ① 75.3 ① 76.0 (28%) ①
FINGERPRINT 80.6 80.4 83.1 ①② 83.3 (36%) ①②
AIDS 97.1 98.3 ① 98.2 ① 98.2 (72%) ①
WEBGRAPHS 80.4 82.3 ① 84.0 ①② 83.2 (15%) ①
Z-test for testing statistical significance (α = 0.05):
①/②/③ Statistically significant improvement over the first/second/third reference system.
ℝ
n
G
F
1
ℝ
n'
φ
Linear PCA
Non-linear kPCA
Ω
Linear SVM
F
2
Non-linear RBF SVM
Fig.1. PCA and Kernel PCA based classification.
non-linear part, and the main difference is the order in which these components are
applied.
Apparently, one could use a combination of kernel PCA based transformation and
non-linear SVM classification. With this setting, however, poor classification results
are achieved on all datasets. We ascribe this performance degradation to overfitting, i.e.
the complexity of the model becomes too high, such that the generalization property is
crucially compromised.
5 Conclusions
The present paper considers the task of classification where graphs are used as rep-
resentation formalism for the underlying objects. Graphs provide us with a versatile
alternative compared to feature vectors, but they suffer from the lack of mathematical
operations in the domain of graphs. However, graph kernels, a relatively novel approach
in pattern recognition and related fields, offer an elegant solution to overcome this ma-
jor drawback of graphs. In the work of this paper we make use of the dissimilarity space
embedding graph kernel which maps the graphs explicitly into an n-dimensional vector
space where each dimension represents the graph edit distance to a predefined proto-
type graph. In contrast with previous papers we use the whole set of training patterns
as prototypes and apply kernelized principal component analysis (kPCA) on the result-
ing vectorial description in order to reduce the dimensionality. The main finding of the

experimental evaluation is that kPCA reduction leads to an improvement of the classifi-
cation accuracy compared to all other systems in general. In particular we observe that
the novel approach with kernelized PCA outperforms the former approach with linear
PCA in four out of six cases.
References
1. Duda, R., Hart, P., Stork, D.: Pattern Classification. 2nd edn. Wiley-Interscience (2000)
2. Jain, A., Dubes, R.: Algorithms For Clustering Data. Prentice-Hall, Englewood Cliffs, NJ
(1988)
3. Jolliffe, I.: Principal Component Analysis. Springer (1986)
4. Conte, D., Foggia, P., Sansone, C., Vento, M.: Thirty years of graph matching in pattern
recognition. Int. Journal of Pattern Recognition and Artificial Intelligence 18(3) (2004) 265–
298
5. G¨artner, T.: A survey of kernels for structured data. SIGKDD Explorations 5(1) (2003)
49–58
6. Riesen, K., Neuhaus, M., Bunke, H.: Graph embedding in vector spaces by means of pro-
totype selection. In Escolano, F., Vento, M., eds.: Proc. 6th Int. Workshop on Graph Based
Representations in Pattern Recognition. LNCS 4538 (2007) 383–393
7. Riesen, K., Kilchherr, V., Bunke, H.: Reducing the dimensionality of vector space embed-
dings of graphs. In Perner, P., ed.: Proc. 5th Int. Conf. on Machine Learning and Data Mining.
LNAI 4571, Springer (2007) 563–573
8. Neuhaus, M., Bunke, H.: Bridging the Gap Between Graph Edit Distance and Kernel Ma-
chines. World Scientific (2007)
9. Bunke, H., Allermann, G.: Inexact graph matching for structural pattern recognition. Pattern
Recognition Letters 1 (1983) 245–253
10. Riesen, K., Neuhaus, M., Bunke, H.: Bipartite graph matching for computing the edit dis-
tance of graphs. In Escolano, F., Vento, M., eds.: Proc. 6th Int. Workshop on Graph Based
Representations in Pattern Recognition. LNCS 4538 (2007) 1–12
11. Wilson, R., Hancock, E., Luo, B.: Pattern vectors from algebraic graph theory. IEEE Trans.
on Pattern Analysis ans Machine Intelligence 27(7) (2005) 1112–1124
12. Wilson, R., Hancock, E.: Levenshtein distance for graph spectral features. In: Proc. 17th Int.
Conf. on Pattern Recognition. Volume 2. (2004) 489–492
13. Luo, B., Wilson, R., Hancock, E.: Spectral embedding of graphs. Pattern Recognition 36(10)
(2003) 2213–2223
14. Duin, R., Pekalska, E.: The Dissimilarity Representations for Pattern Recognition: Founda-
tions and Applications. World Scientific (2005)
15. Bishop, C.: Neural Networks for Pattern Recognition. Oxford University Press (1996)
16. Sch¨olkopf, B., Smola, A., M¨uller, K.R.: Nonlinear component analysis as a kernel eigenvalue
problem. Neural Computation 10 (1998) 1299–1319
17. Shawe-Taylor, J., Cristianini, N.: Kernel Methods for Pattern Analysis. Cambridge Univer-
sity Press (2004)
18. Sch¨olkopf, B., Smola, A.: Learning with Kernels. MIT Press (2002)
19. Bianchini, M., Gori, M., Sarti, L., Scarselli, F.: Recursive processing of cyclic graphs. IEEE
Transactions on Neural Networks 17(1) (2006) 10–18
20. Dosch, P., Valveny, E.: Report on the second symbol recognition contest. In Wenyin, L.,
Llad´os, J., eds.: Graphics Recognition. Ten years review and future perspectives. Proc. 6th
Int. Workshop on Graphics Recognition (GREC’05). LNCS 3926, Springer (2005) 381–397

21. Watson, C., Wilson, C.: NIST Special Database 4, Fingerprint Database. National Institute
of Standards and Technology. (1992)
22. DTP, D.T.P.: AIDS antiviral screen (2004) http://dtp.nci.nih.gov/docs/aids/
aids data.html.
23. Schenker, A., Bunke, H., Last, M., Kandel, A.: Graph-Theoretic Techniques for Web Content
Mining. World Scientific (2005)
