
PORTING LEGACY APPLICATIONS TO IMMERSIVE VIRTUAL
ENVIRONMENTS
A Case Study
Kenny Gruchalla, Jonathan Marbach
Department of Computer Science, University of Colorado at Boulder, USA
Mark Dubin
Department of Molecular,Cellular, and Developmental Biology, University of Colorado at Boulder, USA
Keywords: Immersive visualization, molecular visualization, virtual reality, software engineering, biology.
Abstract: Immersive virtual environments are becoming increasingly common, driving the need to develop new
software or adapt existing software to these environments. We discuss some of the issues and limitations of
porting an existing molecular graphics system, PyMOL, into an immersive virtual environment. Presenting
macromolecules inside an interactive immersive virtual environment may provide unique insights into
molecular structure and improve the rational design of drugs that target a specific molecule. PyMOL was
successfully extended to render molecular structures immersively; however, elements of the legacy
interactive design did not scale well into three-dimensions. Achieving an interactive frame rate for large
macromolecules was also an issue. The immersive system was developed and evaluated on both a shared-
memory parallel machine and a commodity cluster.
1 INTRODUCTION
As immersive environments become more
ubiquitous, so does the desire to adapt legacy three-
dimensional applications to operate within these
environments. However, tightly integrating the
functionality, particularly interactive functionality,
of a legacy three-dimensional application within an
immersive virtual environment (IVE) may not be
possible without major redesign of the application's
architecture. This paper addresses design issues,
implementation issues, and the overall effectiveness
of extending PyMOL (DeLano 2002), an existing
open-source molecular graphics software system, to
support the visualization and manipulation of virtual
macromolecules within an IVE.
PyMOL was successfully extended using
CAVELib
TM
to render its molecule representations
inside a CAVE-like IVE. This allows
crystallographers to physically walk around the
macromolecule representations and view them from
multiple perspectives. Extending the rendering
capabilities was straight-forward; however,
integrating the interactive molecular editing features
of PyMOL was less successful. The initial intent was
to provide a true three-dimensional interface for all
the molecular editing capabilities of PyMOL.
Providing this interface was hindered by two
underlying design principles of PyMOL. First,
virtual objects are selected in PyMOL using an off-
screen buffer color-coding scheme, which is
inappropriate for immersive environments.
Secondly, all PyMOL editing is done directly to the
chemical data structures and not to their graphic
representations. For large molecular structures this
introduces an inefficiency that makes interactive
editing impossible. Therefore, our port is currently
incomplete: it only allows the manipulation
(translation and rotation) of complete molecular
structures but not individual atoms and bonds.
However, this is still a useful result, allowing
crystallographers to investigate the fitting between
two or more molecules.
This work is motivated by a larger pilot study to
determine if there is added value in using immersive
visualization as a molecular research tool. The shape
179
Gruchalla K., Marbach J. and Dubin M. (2007).
PORTING LEGACY APPLICATIONS TO IMMERSIVE VIRTUAL ENVIRONMENTS - A Case Study.
In Proceedings of the Second International Conference on Computer Graphics Theory and Applications - AS/IE, pages 179-184
DOI: 10.5220/0002078401790184
Copyright
c
SciTePress

of a molecule provides the basis for its function.
This is the principle behind rational drug design,
which uses structural information to design drugs
that bind with high affinity and specificity to a
therapeutic target. The effective visualization of
three-dimensional molecular structure can play a
critical role in understanding molecular function,
leading to better drug design. However,
understanding the complex irregular geometry of
multiple macromolecules is a daunting task. Even
more complicated is interacting with and
manipulating these complex three-dimensional
structures. Therefore, immersive visualization of
molecule structures has long been proposed to cope
with the complex three-dimensional nature of
molecule biology (Ihlenfeldt 1997).
There exists a common intuition that an
immersive virtual environment can provide an
improved interface to view and interact with
complex three-dimensional datasets compared to the
more traditional graphics workstation (van Dam
2000). This intuition is based on the fact that an IVE
provides the user with an egocentric three-
dimensional point-of-view, allowing the user to view
and interact with the data space using “natural
skills.” There have been recent studies that have
shown that some three-dimensional tasks can be
improved when the data is presented and
manipulated in an IVE (Pausch 1997, Arns et al.
1999, Swan et al. 2003, Gruchalla 2004). However,
the results are mixed. Several of these and other
studies have also shown reduced performance in
accomplishing various tasks in an IVE (Pausch
1997, Arns et al 1999). The suggestion is that the
applicability of an IVE is highly task dependent.
Therefore, the principle thesis of our larger effort is
to determine if an immersive interface can improve
the rational design of drugs that target a specific
molecule.
2 PYMOL
PyMOL is a powerful and versatile open-source,
cross-platform molecular graphics system with an
embedded Python interpreter. It is capable of real-
time visualization using OpenGL
®
and the
generation of high-quality, ray-traced images.
PyMOL supports most of the common
representations for molecular structures: wire bonds,
cylinders, spheres, ball-and-stick, dot surfaces, solid
surfaces, wire meshes, backbone ribbons, and
cartoon ribbons. Proteins and electron densities can
be imported from many file types (e.g., PDB, SDF,
and MOL) (DeLano 2002).
Figure 1: A PyMOL rendered visualization of a
membrane-bound protein docking to a membrane
(yellow).
PyMOL is written in C, but its primary interface
is an embedded Python interpreter. The basis for its
sophistication is the PyMOL scripting language, a
superset of Python. All of the features of PyMOL
are accessible through a command-line interface.
PyMOL also provides an external Tcl/Tk GUI and a
simple internal OpenGL menu system. Both are
merely wrappers around PyMOL scripting
commands: button presses in the GUI generate
PyMOL script that is then evaluated by the
embedded interpreter (DeLano 2002).
There are several immersive molecular
visualization tools that have already been developed.
VRMol is an immersive application that allows for
the interactive investigation of multiple complex
molecules (Hasse et al. 1996). RealMOL is another
more recent example of an interactive immersive
molecular research tool (Ai and Fröhlich 1998).
However, in the context of our larger research goals,
extending PyMOL provided several advantages over
existing immersive molecular applications. The
driving motivation behind this work is to design an
empirical experiment to study the benefits of using
immersive visualization in rational drug design. A
team of crystallographers, from which we will draw
our test subject population, recommended PyMOL
for its sophistication, quality of visualizations, and
user-adjustable parameters. PyMOL is currently
being used within this target population as a desktop
molecular research tool. Drawing from an
experienced user-base will allow us to conduct a
comparative study of a real-world application on
real-world problems. A secondary factor in the
selection of PyMOL was that it has been released
GRAPP 2007 - International Conference on Computer Graphics Theory and Applications
180
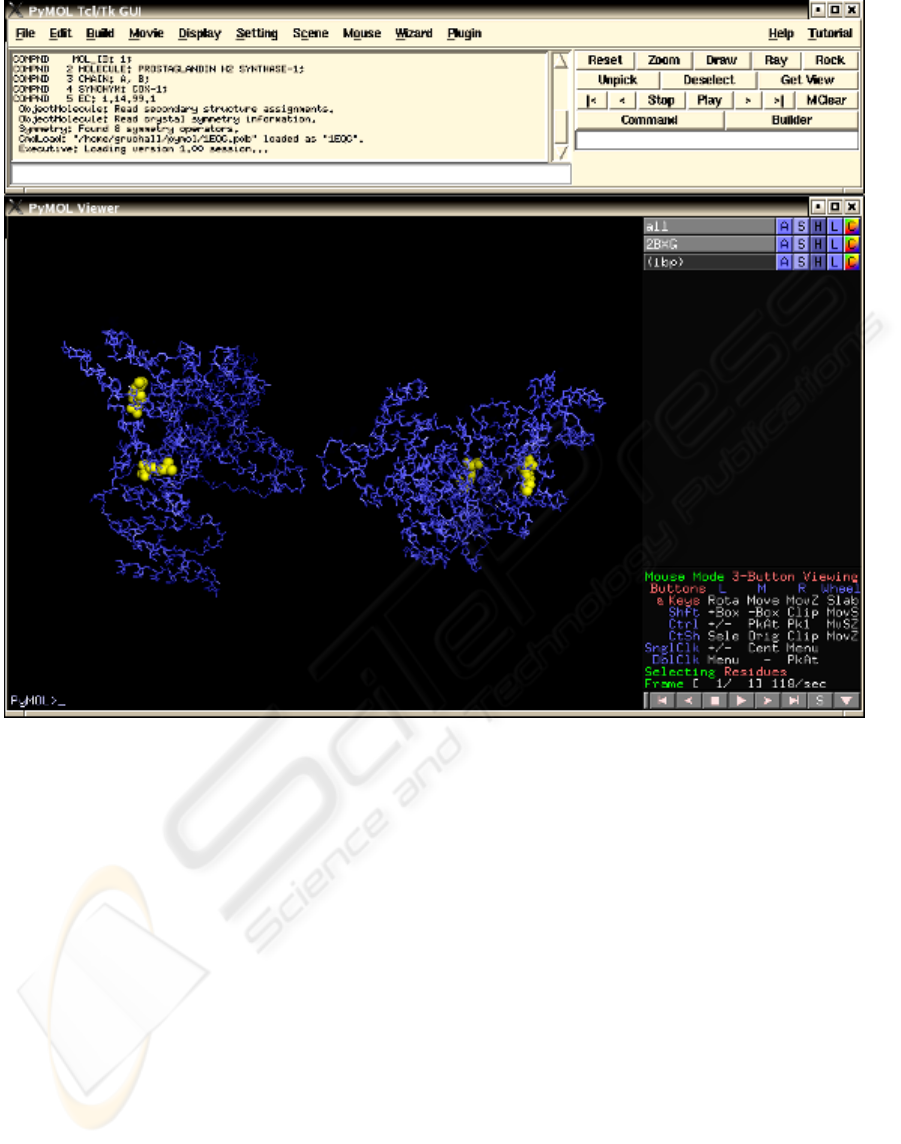
Figure 2: Desktop version of PyMOL (DeLano 2002). PyMOL provides a Python-
b
ased scripting language, which can be
used directly at the command prompt or indirectly through the Tcl/Tk GUI or the simple OpenGL menu system.
under an open-source software license. This
provides us the opportunity to contribute to both the
immersive visualization and molecular research
communities.
3 IMMERSIVE PORT
The intent of the project was to extend PyMOL's
three-dimensional rendering and manipulation
capabilities into an IVE, allowing crystallographers
an egocentric view of molecular structures.
Furthermore, the scientist would be able to
manipulate the macromolecule at a component level:
selecting and manipulating atoms, bonds, and
structures directly using a six-degree of freedom
input device. The goal of the port was not to replace
the desktop version of PyMOL, but rather to
augment and enhance the desktop version with an
immersive counter-part. Three architectural areas of
PyMOL had to be addressed to provide this
functionality: the rendering architecture, thread
control architecture, and interaction architecture.
PyMOL provides two mechanisms to render
molecular structures: the standard OpenGL pipeline
for interactive visualizations, and a built-in ray-
tracer for generating static, high-quality images for
publication. For the purposes of this port, we were
only interested in interfacing with PyMOL's
interactive visualization capabilities. We used
CAVELib
TM
to handle the hardware-specific details
of the IVE display system. An immersive display
callback function was added to PyMOL that is called
from the CAVELib
TM
display loop for each rendered
view. CAVELib
TM
handles the necessary projections
for each view and synchronization among views in
the IVE. The immersive display function is almost
identical to the PyMOL display function, with two
exceptions: it allows CAVELib
TM
to handle
PORTING LEGACY APPLICATIONS TO IMMERSIVE VIRTUAL ENVIRONMENTS - A Case Study
181

stereoscopic rendering, and it bypasses the
additional back-buffer render used in the picking
scheme discussed below.
PyMOL is a multi-threaded application with
separate threads for the rendering and interaction,
the embedded Python interpreter, and the Tcl/Tk
GUI. This architecture was easily extended to handle
the multiple threads used to render each immersive
view. This involved simply adding an additional
mutex to prevent resource competition between the
immersive displays and the windowed display, and
between the immersive interactions and the
command driven interactions. Although the
development required to extend the thread
architecture was minimal, the difficulty of
understanding the PyMOL thread control should not
be understated. Thorough understanding of the
PyMOL and CAVELib
TM
threads was critical in the
success of the port.
PyMOL provides limited but functional
molecular editing capabilities. From the command-
line, entire molecular structures can be manipulated
(e.g., rotated, translated). PyMOL also provides
limited mouse-driven editing capabilities. These
capabilities allow the individual atoms and bonds of
a protein to be selected and transformed. Extending
these capabilities to a true three-dimensional
interface was hindered by two underlying design
principles of PyMOL. First, virtual objects are
selected in PyMOL using a back-buffer color-coding
scheme. When the user clicks a mouse button, a
function renders each pickable object into the back-
buffer using a distinct color. The color at the mouse
position is read back, and is used to index an array
of visible objects. Unfortunately, this process does
not scale to a three-dimensional pointing device. The
position of the three-dimensional pointing device
cannot be projected into the two-dimensions of a
single back-buffer since there may be multiple views
in an immersive environment. This is further
complicated by stereographic projection, since there
may be multiple back-buffers per view. Extending
PyMOL's architecture to support data structures that
can be efficiently selected in three-dimensions will
require a sizable effort. As a temporary work-
around, we have provided the ability to select
objects from the command-line that can then be
manipulated within the IVE.
The second problem in adapting interactions into
three dimensions was the design of the data
structures used to model the molecular structures.
PyMOL editing is done directly to the chemical data
structures and not to their graphic representations:
transformations are applied to each individual atom,
then, the graphical representation of the molecule is
reconstructed from the transformed atoms. For large
molecules, the construction or reconstruction of
certain types of graphics representations (e.g., solid
surfaces) can cause a delay of several seconds. Since
most PyMOL interaction originates from the
command-line, this inefficiency was tolerable.
However, for manipulation of objects inside an IVE
the potential multiple-second lag between the user's
movements and the visualization response was
clearly unacceptable as operational feedback.
Indeed, if the internal structure of a molecule is
modified, the graphics representation may need to be
reconstructed. However, if the internal structure of a
molecule is not edited and only the molecule's
relative position or orientation has been changed, the
graphics representation does not have to be
reconstructed -- it can simply be transformed with a
transformation matrix.
Figure 3: User interactively docking a protein to a
membrane (yellow) inside an immersive virtual
environment.
To allow rapid independent manipulation of
complete molecular structures, we added
transformation matrices to encapsulate the position
and orientation of the graphics representations
during editing. This provides a crystallographer
means to investigate the fitting between two or more
molecules inside the IVE interactively (see Figure
3). Once the relative positions and orientations of the
graphical representations are decided upon, a
scripting command has been added to transform and
synchronize the chemical data structures with the
graphics representations. Currently, there are no
means to interactively modify internal molecular
structures from within the IVE.
GRAPP 2007 - International Conference on Computer Graphics Theory and Applications
182

4 RESULTS
System performance is critical, as our pilot users are
interested in visualizing and interacting with
macromolecules on the order of tens of thousands of
atoms. For example, the macromolecule 1GRU has
over 58,000 atoms (Ranson et al. 2001). Even a
medium-resolution surface representation of this
macromolecule has close to one million triangles. To
maintain satisfactory spatiotemporal correlation
between a user’s actions and system responses, an
update rate of at least ten frames per second must be
achieved (Bowman et al. 2005). Furthermore, the
application is stereoscopic, requiring two views to
be rendered per frame. Therefore, we would require
a throughput of twenty million Gouraud-shaded
polygons with specular highlighting per second. This
throughput figure is of course for one molecule only,
yet for the application to be of value to users, they
must be able to interactively manipulate multiple
molecules.
Our initial development efforts were carried out
on a 20-processor SGI Onyx 3800 with four
InfiniteReality3 graphics pipes. The appeal of
developing on such a machine is the simplicity of
software development: a single application is
executed on a single machine, with multiple
rendering threads running in parallel. The distinct
disadvantage of such a hardware environment is that
its graphics performance trails that of recent
hardware.
To deliver the performance necessary to maintain
interactivity, we ported our immersive version of
PyMOL to a four-PC cluster equipped with
QuadroFX 3000G graphics cards from NVidia. The
clear advantage of such a system is its high-
performance graphics capabilities and cost-effective
upgrade path, but this system incurs its own costs.
With four applications running simultaneously on
separate machines, synchronization becomes an
issue. CAVELib™ does provide synchronized
tracker and controller data to the application;
therefore, if the application responds
deterministically to these inputs, the cluster nodes
will behave identically. Unfortunately, there is no
mechanism that automatically synchronizes
application-specific events and data.
To avoid requiring that the user replicate
commands manually across the four PCs, we have
implemented a command distribution mechanism
using CAVELib™’s synchronized communication
functions. As described earlier, most of PyMOL’s
functionality is exposed through extensions to the
embedded Python interpreter, which accepts
commands as character strings. To achieve
command synchronization, we intercept the strings
before they are sent to the interpreter, and distribute
them to the remote cluster nodes. The only limitation
of this approach is that commands must be entered
on the master node, since CAVELib™ only exposes
a synchronized scatter operation from master to non-
master nodes. A major benefit of this approach is
that most user-initiated events, even from PyMOL
GUIs, are handled as command strings, so these are
automatically distributed and synchronized across
the cluster.
We performed a small pilot study to assess the
usability of the immersive port. Three University of
Colorado biochemistry groups were invited to study
a molecule of their choice using the immersive
version of PyMOL. The biochemistry groups
conducted actual research on how the structure of
their molecule related to its function. Typically,
three or four members of the team would work
collaboratively inside the IVE, while one team
member would control the visualization from a
desktop computer using the PyMOL command-line
and desktop interfaces.
Despite the limitations of the immersive port, our
pilot users indicate that the immersive version of
PyMOL is indeed useful as a research tool. In fact,
the immersive examination of the selected molecules
led all three research groups to a new understanding
of their molecule's functional structure. All three
groups reported the discovery of a large spatial
feature, such as an empty space or ridge that had not
been previously recognized during extensive
previous work with the molecule using the PyMOL
on the desktop.
5 CONCLUSIONS
In this paper, we discussed some of the issues and
limitations of porting PyMOL into an immersive
virtual environment. We successfully extended
PyMOL to render its molecule representations inside
an IVE. However, the interactive molecular editing
features of PyMOL have not been completely nor
effectively integrated into our immersive version. As
a consequence of how three-dimensional objects are
represented and how they are selected in PyMOL,
full immersive integration of the interactive features
will not be realized without a significant amount of
work. This port serves as an example that legacy
three-dimensional applications designed for a
desktop may have made underlying assumptions that
will complicate an immersive port. In fact, to fully
PORTING LEGACY APPLICATIONS TO IMMERSIVE VIRTUAL ENVIRONMENTS - A Case Study
183

integrate the interactive features of a legacy
application may require major architectural changes.
The following observations can be made from the
effort to port PyMOL:
• PyMOL's use of the OpenGL rendering
pipeline provided a direct means to extend
the molecular visualizations to an
immersive platform using CAVELib
TM
.
• Thread-safety was a key issue. Without
PyMOL's existing thread-safe architecture
the immersive extension would have been
vastly more complicated.
• Being based on an embedded Python
interpreter, commands strings were easily
distributed across a cluster.
• Some object selection schemes designed for
two-dimensional pointing devices do not
scale well for three dimensional pointing
devices.
• Although PyMOL provided molecular
editing capabilities, these were not
designed to support the interactivity
necessary in an IVE.
Much work remains before the immersive PyMOL
will be fully capable of supporting rational drug
design. The ability to select molecules, bonds, and
atoms using a three-dimensional pointing device will
be critical to the success of the project. New data
structures will need to be added to PyMOL to
support this ability. Additionally, the performance of
PyMOL must be improved to reach the frame rates
necessary for interaction with large macromolecules.
Since PyMOL was not architected for rendering to
multiple windows, all OpenGL rendering must be
done in immediate mode. We are currently
investigating options for improving performance,
such as allowing display lists to be used in the
presence of multiple rendering contexts. Finally,
since PyMOL's internal GUIs are rendered using
OpenGL, we have begun investigating the
possibility of directly rendering them in the
immersive environment. Although we do not aim to
replicate all of PyMOL's functionality within the
IVE, this should improve the overall usability of the
immersive port.
ACKNOWLEDGEMENTS
This project was supported by a University of
Colorado Butcher Award and by equipment
donations from NVIDIA. We would like to thank
Geoffery Dorn, Gwen Pech, and Mick Coady of the
University of Colorado at Boulder, BP Center for
Visualization for their assistance, support and
advice. We are most grateful to the members of the
research groups who participated in the pilot study.
Professor Pardi was especially helpful in defining
the early stages of this project.
REFERENCES
Ai, Z. and Frohlich, T., 1998. Molecular Dynamics
Simulation in Virtual Environments. Computer
Graphics Forum. 17(3), 267-275.
Arns, L., Cook, D., Cruz-Neira, C., 1999. The benefits of
statistical visualization in an immersive virtual
environment. In Proceedings of IEEE Virtual Reality
1999. IEEE Press, 88-95.
Bowman, D.A., Kruijff, E., LaViola, J., Poupyrev, I.,
2005. 3D user Interfaces: Theory and Design.
Addison-Wesley.
DeLano, W.L., 2002. The PyMOL Molecular Graphics
System. DeLano Scientific, San Carlos, CA, USA.
http://www.pymol.org
Gruchalla, K., 2004. Immersive Well-Path Editing:
Investigating the added value of immersion. In
Proceedings of IEEE Virtual Reality 2004, IEEE
Press, 157-164.
Haase, H., Strassner, J., and Dai, F., 1996. VR techniques
for the investigation of molecule data. Computers &
Graphics, 20(2), 1996, 207-217.
Ihlenfeldt, W., 1997. Virtual Reality in Chemistry. Journal
of Molecular Modeling 3, 386-402.
Moshell, J.M., and Hughes, C.E., 2002. Virtual
environments as a tool for academic learning. In:
Stanney, K.M., (ed.) Handbook of Virtual
Environments. Lawrence Erlbaum Associates:
Mahwah, NJ. Chapter 45. pp 893-910.
Pausch, R., Proffitt, D., Williams, G., 1997. Quantifying
Immersion in Virtual Reality. Proceedings of the 24th
annual conference on computer graphics and
interactive techniques, 13-18.
Ranson, N., Farr, G., Roseman, A., Gowen, B., Fenton,
W., Horwich, A., Saibil, H., 2001. ATP-Bound States
of Groel Captured by Cyro-Electron Microscopy. Cell.
107. 869.
Swan, J., Gabbard, J., Hix, D., Schulman, R., and Kim, K.,
2003. A Comparative Study of User Performance in a
Map-Based Virtual Environment. Proceedings of
IEEE Virtual Reality 2003, IEEE Press, 259-266.
GRAPP 2007 - International Conference on Computer Graphics Theory and Applications
184
