
LOG-UNBIASED LARGE-DEFORMATION IMAGE REGISTRATION
Igor Yanovsky, Stanley Osher
Department of Mathematics, University of California, Los Angeles
405 Hilgard Avenue, Los Angeles, CA 90095-1555, USA
Paul M. Thompson, Alex D. Leow
Laboratory of Neuro Imaging, UCLA School of Medicine
635 Charles E. Young Drive South, Los Angeles, CA 90095-7332, USA
Keywords:
Nonlinear image registration, information theory, mutual information, log-unbiased deformation, biomedical
imaging.
Abstract:
In the past decade, information theory has been studied extensively in medical imaging. In particular, image
matching by maximizing mutual information has been shown to yield good results in multi-modal image
registration. However, there has been few rigorous studies to date that investigate the statistical aspect of the
resulting deformation fields. Different regularization techniques have been proposed, sometimes generating
deformations very different from one another. In this paper, we apply information theory to quantifying the
magnitude of deformations. We examine the statistical distributions of Jacobian maps in the logarithmic
space, and develop a new framework for constructing log-unbiased image registration methods. The proposed
framework yields both theoretically and intuitively correct deformation maps, and is compatible with large-
deformation models. In the results section, we tested the proposed method using pairs of synthetic binary
images, two-dimensional serial MRI images, and three-dimensional serial MRI volumes. We compared our
results to those computed using the viscous fluid registration method, and demonstrated that the proposed
method is advantageous when recovering voxel-wise local tissue change.
1 INTRODUCTION
Non-linear image registration is a well-established
field in medical imaging with many applications
in functional and anatomic brain mapping, image-
guided surgery, and multimodality image fusion
(Avants and Gee, 2004; Grenander and Miller, 1998;
Thompson and Toga, 2002). The goal of image reg-
istration is to align, or spatially normalize, one im-
age to another. In multi-subject studies, this serves
to reduce subject-specific anatomic differences by de-
forming individual images onto a population average
brain template.
The deformations that map each anatomy onto a
common standard space can be analyzed voxel-wise
to make inferences about relative volume differences
between the individuals and the template, or statistical
differences in anatomy between populations. Simi-
larly, in longitudinal studies it is possible to visual-
ize structural brain changes that occur over time by
deforming subjects’ baseline scans onto their subse-
quent scans, and using the deformation map to quan-
tify local changes. This general area of computa-
tional anatomy has become known as tensor-based
morphometry (Davatzikos et al., 1996; Shen and Da-
vatzikos, 2003; Thompson et al., 2000).
To construct a deformation that is one-to-one and
differentiable (Christensen et al., 1996; Miller, 2004;
Holm et al., 2004), we must impose a regularizing
constraint. Thus, the problem of image registration is
often cast as a minimization problem with a combined
cost functional consisting of an image matching func-
tional and a regularizing constraint on the deforma-
tion. Common choices of image matching functional
include squared intensity difference, cross correlation
(Collins et al., 1994), and (normalized) mutual in-
formation or other divergence-based or information-
theoretic measures (D’Agostino et al., 2003; He et al.,
2003; Pluim et al., 2004), while choices of regular-
ization usually involve differential operators inspired
by thin-plate spline theory, elasticity theory, fluid
dynamics and the Euler-Poincare equations (Miller,
272
Yanovsky I., Osher S., M. Thompson P. and D. Leow A. (2007).
LOG-UNBIASED LARGE-DEFORMATION IMAGE REGISTRATION.
In Proceedings of the Second International Conference on Computer Vision Theory and Applications - IFP/IA, pages 272-279
Copyright
c
SciTePress

2004; Thompson and Toga, 2000).
2 THEORY
2.1 Global Preservation of Density
Maps
In this paper, we study smooth deformations
~
h that
map computational domain Ω bijectively onto itself.
Let us assume, without loss of generality, that the vol-
ume of this domain is 1, i.e., |Ω| = 1. The inverse map
of
~
h is denoted as
~
h
−1
and the Jacobian matrix of
~
h as
D
~
h. The Jacobian map can thus be defined as the de-
terminant of the Jacobian matrix |D
~
h|.
In volumetric studies, the determinant of the Jaco-
bian matrix (density) applied to any given deforma-
tion
~
h is an important quantity, encoding the voxel-
wise volume change. As
~
h (and
~
h
−1
) is bijective and
thus globally volume preserving, we have the follow-
ing preservation of global density:
Ω
|D
~
h(ξ)|dξ =
Ω
d~y = 1,
Ω
|D
~
h
−1
(ξ)|dξ =
Ω
d~x = 1.
(1)
Given global preservation of density maps, we can as-
sociate three probability density functions to
~
h,
~
h
−1
,
and the identity map (id):
P
h
(·) = |D
~
h(·)|,
P
h
−1
(·) = |D
~
h
−1
(·)|,
P
id
(·) = 1.
(2)
Differentiating the identity
~
h
−1
(
~
h(~x)) = ~x on both
sides and setting~y =
~
h(~x), we obtain
D
~
h
−1
(~y) · D
~
h(~x) = id, (3)
and hence,
|D
~
h
−1
(~y)| · |D
~
h(~x)| = 1. (4)
By identifying deformations with their corre-
sponding global density maps, we can now apply in-
formation theory to quantifying the magnitude of de-
formations. In our approach, we choose the symmet-
ric Kullback-Leibler (sKL) distance:
sKL(P
h
,P
id
) = KL(P
id
,P
h
) + KL(P
h
,P
id
) (5)
to measure the magnitude of any deformation
~
h. Here
KL, the Kullback-Leibler distance between two prob-
ability density functions X and Y, is defined as
KL(X,Y) =
Ω
X log
X
Y
d~x ≥ 0. (6)
To motivate this approach, notice that the first part
of sKL measure is simply integrating the log-density
over the entire computational image domain:
Ω
log|D
~
h(~x)|d~x = −
Ω
log
1
|D
~
h(~x)|
d~x
= −
Ω
P
id
log
P
id
P
h
d~x
= −KL(P
id
,P
h
) ≤ 0.
(7)
To attach geometric meaning to the second term, we
notice that the KL distance has skew-symmetry with
respect to
~
h and its inverse
KL(P
id
,P
h
−1
) = −
Ω
log|D
~
h
−1
(~y)|d~y
=
Ω
log|D
~
h(~x)|
|D
~
h(~x)|d~x
=
Ω
P
h
log
P
h
P
id
d~x
= KL(P
h
,P
id
),
(8)
where the second equality was obtained using a
change of variables,~y =
~
h(~x). Similarly, we have
KL(P
id
,P
h
) = KL(P
h
−1
,P
id
). (9)
2.2 Unbiased Deformation in the
Logarithmic Space
Before developing formulations to construct unbiased
deformations in the logarithmic space, we generalize
equation (7) to the case of mapping regions of interest
(ROI). Assuming we have a priori knowledge that one
ROI is mapped to another, we would like to recover a
mapping that is unbiased in the logarithmic space. In-
tuitively, without further knowledgeother than overall
ROI matching, the resulting Jacobian map should take
a constant value inside the ROI. This can be achieved
using the proposed formulations. Indeed, given any
deformation~g mapping domain A in the source (with
volume a) to domain B in the target (with volume b),
we have the following
1
a
A
log|D~g(~x)|d~x ≤ log
b
a
, (10)
with equality obtained if and only if the Jacobian map
of~g takes a constant value (i.e., b/a). This generaliza-
tion can be shown by observing that the logarithmic
mapping is a convex mapping:
∑
n
log(x
i
) ≤ nlog(¯x); ¯x =
1
n
∑
n
x
i
. (11)
With the above generalization, one can see that, as-
suming the only constraint being an ROI deformation

(a) (b)
(a) (b)
(c) (d)
Figure 1: Circle-to-Ellipse example. (a) image T; (b) image
S; (c) image T is deformed to image S using Christensen’s
model; (d) image T is deformed to image S using the pro-
posed model. Blue, yellow and red contours represent the
boundaries of objects in T, S, and deformed T, respectively.
Note that for both methods, yellow contour is essentially
invisible due to a very close match. However, the resulting
grid of the proposed method is visually more regular.
(a) (b)
(a) (b)
Figure 2: Circle-to-Ellipse example. Jacobian map of the
deformation using (a) Christensen’s model and (b) the pro-
posed model.
from A to B, the unbiased mapping under the logarith-
mic operation has an evenly distributed Jacobian field,
which is also intuitively correct (as there is no reason
to assume non-uniformity of the Jacobian field).
Given equation (7) and its generalization, we now
propose to quantify the distance between any given
deformation and the identity map by computing the
symmetric KL distance through their density func-
tions. Due to the above mentioned skew-symmetry,
this distance takes the following several equivalent
0 0.2 0.4 0.6 0.8 1 1.2
0
200
400
600
800
1000
Jacobian
Histogram
Christensen‘s model
0 0.2 0.4 0.6 0.8 1 1.2
0
200
400
600
800
1000
Jacobian
Histogram
proposed model
Figure 3: Circle-to-Ellipse example. Histograms of Jaco-
bian values of the deformations inside the ellipse for Chris-
tensen’s model and the proposed model.
0 50 100 150 200 250 300 350 400 450 500
0
0.02
0.04
0.06
0.08
0.1
0.12
0.14
0.16
0.18
Iteration number
Standard deviation
Circle−to−Ellipse example: The standard deviation of Jacobian values
Christensen‘s model
proposed model
(a)
0 50 100 150 200 250 300 350 400 450 500
0
0.02
0.04
0.06
0.08
0.1
0.12
0.14
0.16
0.18
Iteration number
Symmetric KL distance
Circle−to−Ellipse example: The symmetric KL distance
Christensen‘s model
proposed model
(b)
Figure 4: Circle-to-Ellipse example. (a) Standard deviation
of Jacobian values inside the ellipse per iteration. (b) Sym-
metric KL distance. For Christensen’s model (dashed blue),
both standard deviation and symmetric KL distance increase
while for the proposed model (solid red), both standard de-
viation and symmetric KL distance stabilize.
forms:
sKL(P
h
,P
id
) = sKL(P
h
−1
,P
id
)
= KL(P
h
,P
id
) + KL(P
h
−1
,P
id
)
= KL(P
h
,P
id
) + KL(P
id
,P
h
)
= KL(P
id
,P
h
−1
) + KL(P
id
,P
h
)
= KL(P
id
,P
h
−1
) + KL(P
h
−1
,P
id
)
=
Ω
|D
~
h(~x)| − 1
log|D
~
h(~x)|d~x
=
Ω
|D
~
h
−1
(~y)| − 1
log|D
~
h
−1
(~y)|d~y.
(12)
To see why minimizing equation (12) leads to unbi-
ased deformation in the logarithmic space, we ob-
(a) (b)
(a) (b)
(c) (d)
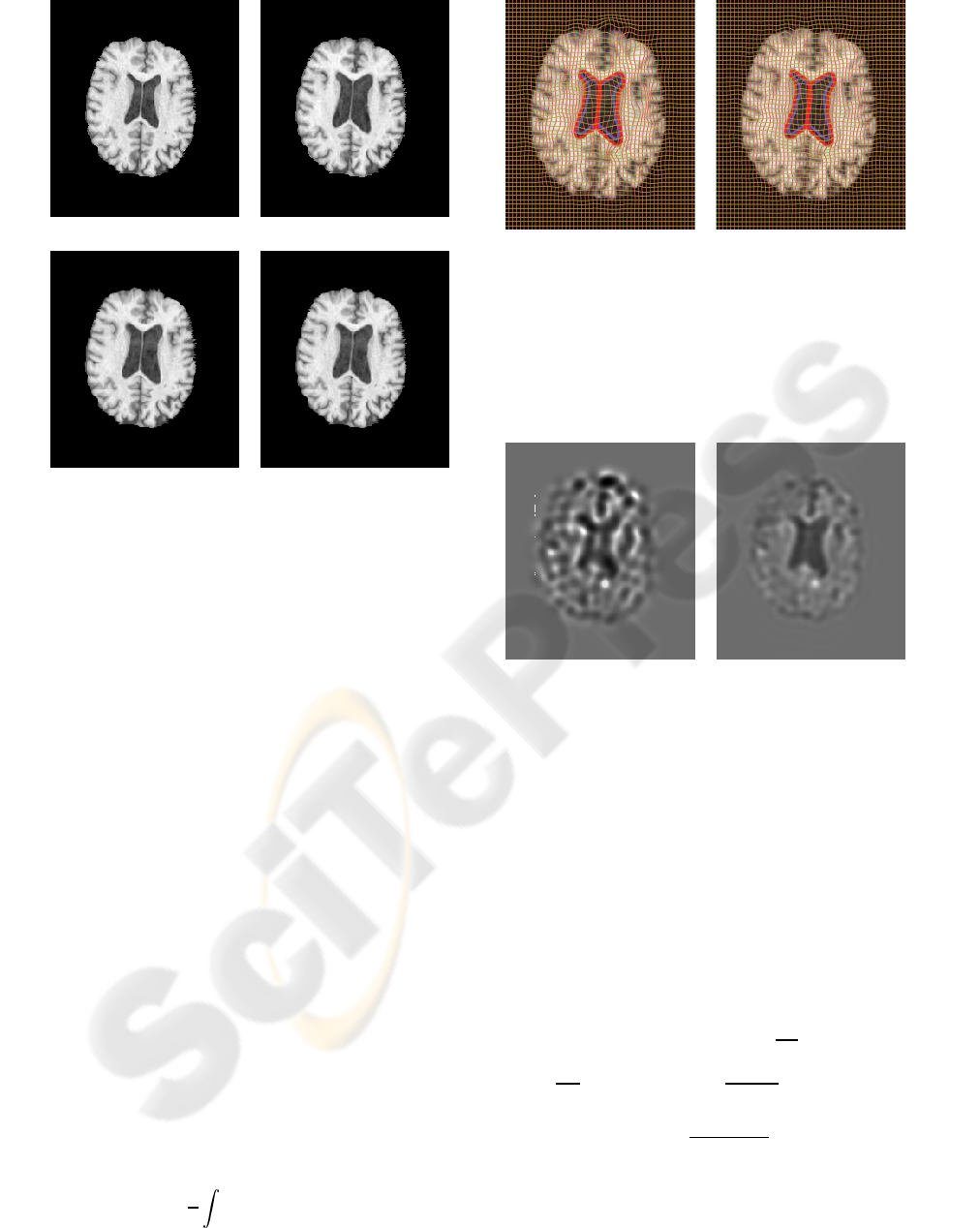
Figure 5: Serial MRI example. (a) image T; (b) image S; (c)
image T is deformed to image S using Christensen’s model;
(d) image T is deformed to image S using the proposed
model.
serve that the integrand is always non-negative, and
only evaluates to zero when
~
h is volume-preserving
everywhere (Jacobian of
~
h is 1 everywhere). Thus,
by treating it as a cost, we recover zero-change by
minimizing this cost when we compare images dif-
fering only in noise. Also, this approach is unbiased
for mapping ROIs in the logarithmic space, due to the
inequality in (10).
3 IMPLEMENTATION
Let us denote the template image as T(~x) and the
study image as S(~x) defined on the spatial domain Ω.
We solve for deformation
~
h, such that T ◦
~
h matches S,
while minimizing the symmetric KL distance in equa-
tion (12). The deformation
~
h is usually expressed at
each voxel in terms of the displacement vector~u from
the original position:
~
h(~x) = ~x −~u(~x). In this paper,
we will use the sum of the squared differences (SSD)
to measure the accuracy of matching between the de-
formed template and the study:
SSD(T,S,~u) =
1
2
Ω
|T(~x−~u) − S(~x)|
2
d~x, (13)
(a) (b)
(a) (b)
Figure 6: Serial MRI example. Results obtained with (a)
Christensen’s model and (b) the proposed model. Blue, yel-
low and red contours represent the boundaries of ventricles
in T, S, and deformed T, respectively. Note that for both
methods, yellow contour is essentially invisible due to a
very close match. However, the resulting grid of the pro-
posed method is visually more regular.
(a) (b)
(a) (b)
Figure 7: Serial MRI example. Jacobian map of the defor-
mation using (a) Christensen’s model and (b) the proposed
model.
which is also known as a Gaussian sensor model. To
numerically implement our approach, we propose to
minimize a combined cost function
C = SSD+ λ(sKL). (14)
This can be achieved using incremental updating
along the gradient descent of the correspondingEuler-
Lagrange equation. Hence, we obtain the ith compo-
nent of the force field:
f
i
(~x,~u(~x,t)) = −[T(~x−~u) − S(~x)]
∂T
∂x
i
~x−~u
−λ
∑
j
∂
∂x
j
1+ log|D
~
h(~x)| −
1
|D
~
h(~x)|
Co
ij
(~x)
,
D
~
h(~x)
−1
=
Co
ij
(~x)
T
|D
~
h(~x)|
,
(15)
where Co
ij
is the matrix cofactor of the (i, j)-th com-
ponent of the Jacobian matrix D
~
h.
0 0.2 0.4 0.6 0.8 1 1.2 1.4 1.6 1.8
0
100
200
300
400
500
600
Jacobian
Histogram
Christensen‘s model
0 0.2 0.4 0.6 0.8 1 1.2 1.4 1.6 1.8
0
100
200
300
400
500
600
Jacobian
Histogram
proposed model
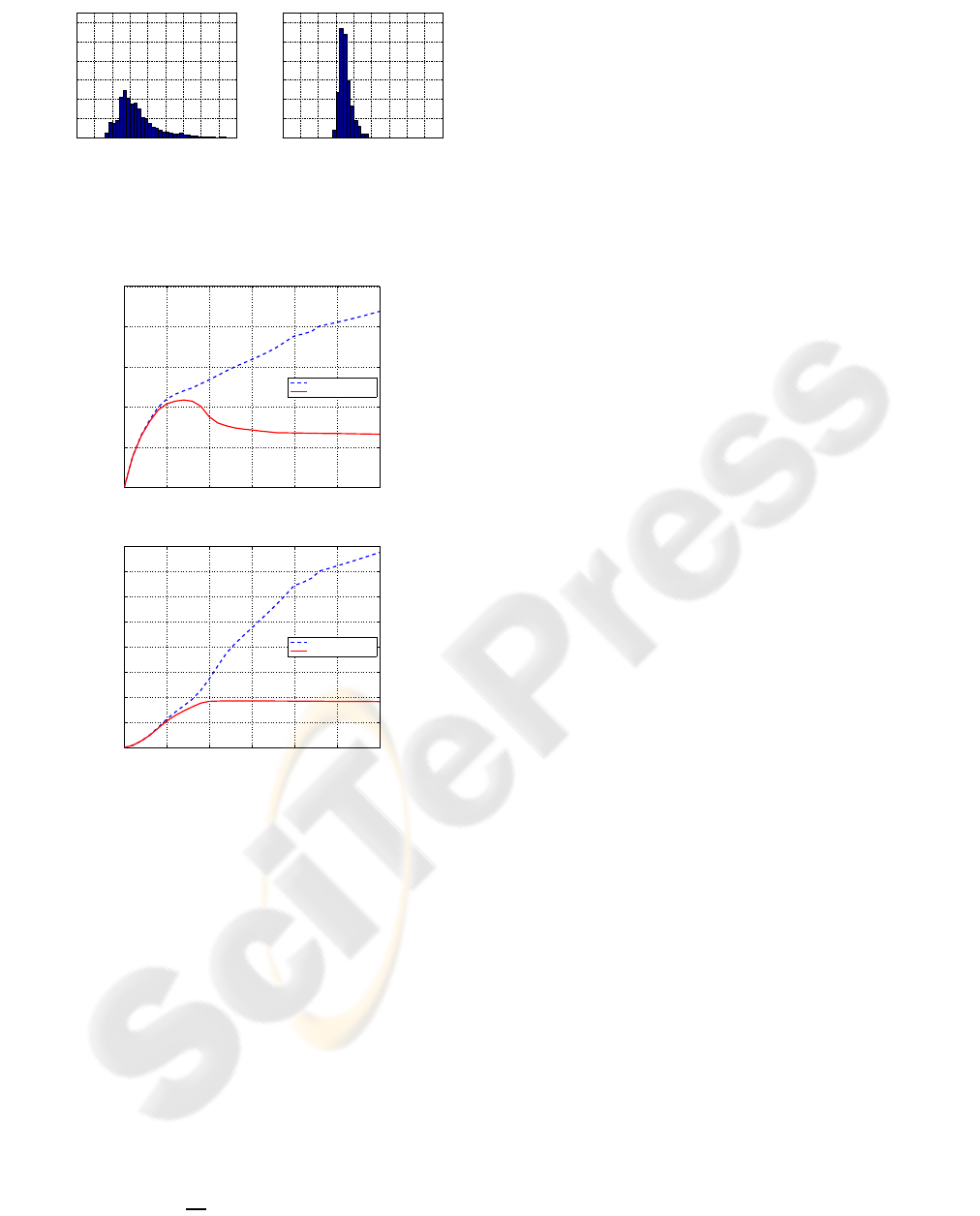
Figure 8: Serial MRI example. Histograms of Jacobian val-
ues of the deformations inside ventricles for Christensen’s
model and the proposed model.
0 50 100 150 200 250 300
0
0.05
0.1
0.15
0.2
0.25
Iteration number
Standard deviation
MRI example: The standard deviation of Jacobian values
Christensen‘s model
proposed model
(a)
0 50 100 150 200 250 300
0
0.005
0.01
0.015
0.02
0.025
0.03
0.035
0.04
Iteration number
Symmetric KL distance
MRI example: The symmetric KL distance
Christensen‘s model
proposed model
(b)
Figure 9: Serial MRI example. (a) Standard deviation of
Jacobian values inside the ventricle per iteration. (b) Sym-
metric KL distance. For Christensen’s model (dashed blue),
both standard deviation and symmetric KL distance increase
while for the proposed model (solid red), both standard de-
viation and symmetric KL distance stabilize.
In this paper, we follow the approach in
(D’Agostino et al., 2003) solving the viscous fluid
model (Christensen et al., 1996). Of note, in (Chris-
tensen et al., 1996), the authors used the sum of the
squared differences (SSD) as a cost functional for
minimization (no control over the distribution of the
Jacobian values was employed). Given the velocity
field~v, the following partial differential equation can
be solved to obtain the displacement field~u:
∂~u
∂t
=~v−~v·
~
∇~u. (16)
The instantaneous velocity as in (D’Agostino et al.,
2003) is obtained by convolving
~
f with Gaussian ker-
nel G
σ
of variance σ:
~v = G
σ
∗ (−
~
f(~x,~u)). (17)
4 RESULTS AND DISCUSSION
In this section, we implemented and tested the pro-
posed nonlinear registration model. The deforma-
tion fields were computed using adaptive time step-
ping, with maximal change in displacement of 0.1 al-
lowed in each iteration. In order to obtain a fair com-
parison between the proposed and the viscous fluid
method, re-gridding was not employed. Re-gridding
is essentially a memoryless procedure, as how im-
ages are matched after each re-gridding is indepen-
dent of the deformation before the re-gridding, ren-
dering the comparison of final Jacobian fields and cost
functionals problematic. Moreover, the strategy of re-
gridding, through the relaxation of deformation over
time, is less rigorous from a theoretical standpoint.
In order to gain more insight into the effect of the
symmetric KL distance term in (12), we first consider
matching two binary synthetic images. In Figures 1
through 4, we show the results of deforming a disk
into an ellipse (both 128 by 128; λ = 500 in (15)).
As seen in Figure 1(c,d), both the fluid registration
(Christensen’s) model and the proposed model gener-
ated a close match between the deformed image and
the study. Here, optimal matching was considered
achieved once the overall cost functional stopped de-
creasing. However, as seen in Figures 2 and 3, the
proposedmethod more evenly distributesdeformation
inside and outside an ellipse (resulting from the con-
vex property of the logarithmic mapping in inequality
(10)). Note the vertical stretching of the grid in the
center of the ellipse for the proposed method, which
is a consequence of uniform distribution of Jacobian
values. On the other hand, using Christensen’s model,
the grid does not uniformly adjust to object’s volume
change; this is especially noticeable in the center of
the ellipse. Figure 4(a) plots the standard deviation
of the Jacobian field inside the ellipse as a function
of iteration number. For Christensen’s model, the
standard deviation inside the ellipse increased with
the number of iterations, while the proposed method
yielded an optimized standard deviation as more it-
erations were computed. The proposed symmetric
KL distance also increased for Christensen’s method,
while it was minimized for the proposed method as
shown in Figure 4(b).
In Figures 5 through 9, we show the results of
matching a pair of 2D slices from a set of Serial MRI
(a) (b) (c) (d)
(a) (b) (c) (d)
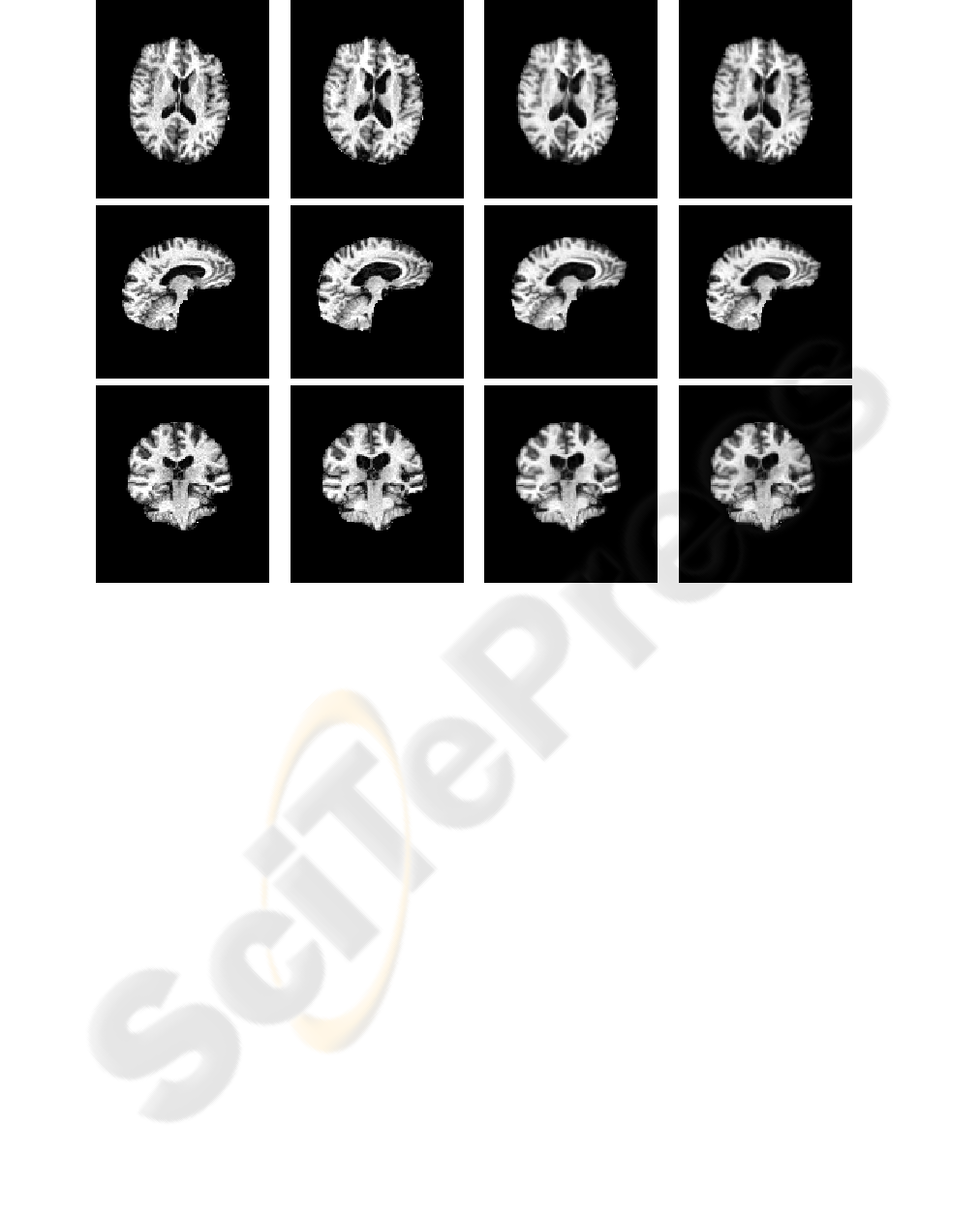
Figure 10: 3D Serial MRI example. Rows depict slices in axial (row 1), sagittal (row 2), and coronal (row 3) planes. Columns
depict (a) T; (b) S; (c) T deformed using Christensen’s model; (d) T deformed using the proposed model.
images (each of size 226 by 256; λ = 400 in (15)),
where visually significant ventricle enlargement is
present. Both Christensen’s method and the proposed
model generated a close match between the deformed
image and the study (Figure 5(a-d)). Here, there is no
reason to not evenly distribute Jacobian field inside
the ventricles, as realized using the proposed method.
In contrast, Christensen’s method generated a density
map with extreme values along the ventricular bound-
ary. Indeed, given the overall longitudinal ventricu-
lar dilatation, we argue that the corresponding density
change map should be constant inside the ventricle.
As seen in Figure 9, both the standard deviation inside
the ventricle and the symmetric KL distance increased
for Christensen’s method, while these quantities sta-
bilized for the proposed method.
In the last numerical example (Figures 10
through 12), we tested the proposed model using
two 3D Serial MRI volumes obtained from a patient
with right-side temporal atrophy (6 years apart; each
of size 112x128x128; λ = 500). In this example,
the same conclusions were reached, demonstrating
both the numerical and theoretical advantages of our
method. In particular, in Figure 11(b), right temporal
atrophy (RT) and ventricular enlargement (V) are eas-
ily visualized in the Jacobian map generated using the
proposed method, while Christensen’s method gener-
ated a very noisy map (Figure 11(a)).
5 FUTURE DIRECTIONS
This paper introduces a new framework for the con-
struction of diffeomorphic maps that yield theoreti-
cally and intuitively correct Jacobian statistics. Simi-
lar concept can be applicable to constructing joint reg-
istration and segmentation algorithms, with the lat-
ter based on Jacobian values. To this end, we are
currently investigating a level set method (Osher and
Sethian, 1988; Osher and Fedkiw, 2003) based imple-
mentation (Chan and Vese, 2001) that would alow us
to simultaneously register serial images and identify
regions of atrophy/expansion.
The idea of employing symmetric KL distance in
nonlinear image registration presented in this work

(a) (b)
Figure 11: 3D Serial MRI example. Jacobian map overlaid
with the deformed volume for Christensen’s model (column
a) and the proposed model (column b), with localized atro-
phy in right temporal area. Rows depict slices in axial (rows
1 and 2), sagittal (row 3), and coronal (row 4) planes.
is also closely related to other scientific fields. For
example, optimization problems involving Jacobian
operator are commonly encountered in grid gener-
ation (Liseikin, 1999) and in continuum mechanics
and computational physics, where the Hencky tensor
arises in modeling very large deformations. However,
we believethat the logarithmic transform has not been
0 100 200 300 400 500 600 700
0
0.05
0.1
0.15
0.2
0.25
Iteration number
Standard deviation
3D MRI example: The standard deviation of Jacobian values
Christensen‘s model
proposed model
(a)
0 100 200 300 400 500 600 700
0
0.002
0.004
0.006
0.008
0.01
0.012
Iteration number
Symmetric KL distance
3D MRI example: The symmetric KL distance
Christensen‘s model
proposed model
(b)
Figure 12: 3D Serial MRI example. (a) Standard devia-
tion of Jacobian values inside the ventricle per iteration. (b)
Symmetric KL distance.
formally introduced in the grid generation literature
and may also be useful there.
ACKNOWLEDGEMENTS
The authors would like to thank James Becker and
Simon Davis at the University of Pittsburgh for pro-
viding the MRI dataset.
This work was supported by Grants U54
RR021813 NIH/NCRR, Grant U01 AG024904,
and Grants R21 RR019771, EB01651, AG016570,
NS049194 to PT.
REFERENCES
Avants, B. and Gee, J. C. (2004). Geodesic estimation for
large deformation anatomical shape averaging and in-
terpolation. NeuroImage, 23. suppl. 1, S139-50.
Chan, T. F. and Vese, L. A. (2001). Active contours with-
out edges. IEEE Transactions on Image Processing,
10(2):266–277.
Christensen, G., Rabbitt, R., and Miller, M. (1996). De-
formable templates using large deformation kine-

matics. IEEE Transactions on Image Processing,
5(10):1435–1447.
Collins, D. L., Peters, T. M., and Evans, A. C. (1994). Auto-
mated 3d nonlinear deformation procedure for deter-
mination of gross morphometric variability in human
brain. In Proceedings of The International Society for
Optical Engineering (SPIE) 2359, pages 180–190.
D’Agostino, E., Maes, F., Vandermeulen, D., and Suetens,
P. (2003). A viscous fluid model for multimodal
non-rigid image registration using mutual informa-
tion. Medical Image Analysis, 7:565–575.
Davatzikos, C., Vaillant, M., Resnick, S. M., Prince, J. L.,
Letovsky, S., and Bryan, R. N. (1996). A computer-
ized approach for morphological analysis of the cor-
pus callosum. J. Comput. Assist. Tomogr., 20(1):88–
97.
Grenander, U. and Miller, M. I. (1998). Computational
anatomy: An emerging discipline. Quarterly of Ap-
plied Mathematics, 56:617–694.
He, Y., Hamza, A. B., and Krim, H. (2003). A general-
ized divergence measure for robust image registration.
IEEE Trans. Signal Process, 51(5). 1211-20.
Holm, D. D., Ratnanather, J. T., Trouve, A., and Younes, L.
(2004). Soliton dynamics in computational anatomy.
NeuroImage, 21. suppl. 1, S170-S178.
Liseikin, V. (1999). Grid Generation Methods. Springer-
Verlag, Heidelberg.
Miller, M. I. (2004). Computational anatomy: shape,
growth, and atrophy comparison via diffeomorphisms.
NeuroImage, 23. suppl. 1, S19-S33.
Osher, S. and Fedkiw, R.(2003). Level Set Methods and Dy-
namic Implicit Surfaces. Applied Mathematical Sci-
ences. Springer-Verlag, New York.
Osher, S. and Sethian, J. (1988). Fronts propagating
with curvature dependent speed; algorithms based
on Hamilton-Jacobi formulations. J. Comput. Phys.,
79:12–49.
Pluim, J. P. W., Maintz, J. B. A., and Viergever, M. A.
(2004). f-information measures in medical image reg-
istration. IEEE Transactions on Medical Imaging,
23(12). 1508-1516.
Shen, D. and Davatzikos, C. (2003). Very high-
resolution morphometry using mass-preserving defor-
mations and hammer elastic registration. NeuroImage,
18(1):28–41.
Thompson, P. M., Giedd, J. N., Woods, R. P., MacDon-
ald, D., Evans, A. C., and Toga, A. W. (2000).
Growth patterns in the developing brain detected by
using continuum mechanical tensor maps. Nature,
404(6774):190–3.
Thompson, P. M. and Toga, A. W. (2000). Elastic image
registration and pathology detection. In Bankman, I.,
editor, Handbook of Medical Image Processing. Aca-
demic Press.
Thompson, P. M. and Toga, A. W. (2002). A framework for
computational anatomy. Computing and Visualization
in Science, 5:13–34.
