
TOWARDS RELIABLE AUTOFOCUSING IN AUTOMATED
MICROSCOPY
Silvie Luisa Br
´
azdilov
´
a
Faculty of Informatics, Masaryk University, Botanick
´
a 68a, Brno, Czech Republic
Keywords:
Fluorescence microscopy, focus function, genetic programming.
Abstract:
The results presented in this paper are twofold. First, autofocusing in automated microscopy is studied and
evaluated with respect to biomedical samples whose images can have more than one in-focus planes. While
the proposed procedure for finding the maximum of a focus function in a short time works satisfactorily, the
focus function itself is identified as the weakest link of the whole process.
Second, an interesting property of functions used for genetic programming, and an algorithm for generating
new individuals are introduced. Their usefulness and applicability are demonstrated on the problem of finding
a new focus function for automated autofocusing in microscopy.
1 INTRODUCTION
An automated microscope system is a powerful tool
for biomedical research, especially in fluorescence
microscopy (Boddeke, 1999). The control of focus is
done through the use of motorized z-axis and an aut-
ofocus algorithm. The algorithm is an iterative pro-
cedure which steps along the z-axis to find the plane
of the best focus. For each z-position an image is ac-
quired from which a focus value is computed accord-
ing to a certain focus function. Actually, the focus
function is a function of the z-position. It says how
sharp the image at the given z-position is. Ideally it
has one clear maximum whose position corresponds
to the z-position of the sharpest image.
According to (Groen et al., 1985) a useful focus
function should fulfil several criteria such as: uni-
modality, accuracy, reproducibility, insensitivity to
other parameters, the extremum be broadly tailed.
Although many focus algorithms have been pro-
posed and compared (Groen et al., 1985; Firestone
et al., 1991; Santos et al., 1997; Geusebroek et al.,
2000; Sun et al., 2004; Bueno-Ibarra et al., 2005), the
selection of an appropriate focus algorithm for spe-
cific conditions remains ad hoc and time consuming.
In fluorescence microscopy weak light signals are
imaged in general and therefore more consideration
is needed in choosing a focus function compared to
bright field microscopy. Due to the placement of the
objects (i. e., tissue or large cells) on the slides, the
thickness of the objects, and possible bending of the
slide, the objects normally do not lie in one plane
and hence the focus function is usually not unimodal.
This can consequently cause problems when search-
ing for the position of the maximum focus, because
many methods, such as Boddeke’s algorithm (Bod-
deke et al., 1994), rely on unimodality of the focus
function.
This paper is based on experiments with auto-
focusing simulated on a personal computer in the
MATLAB environment. The results of these exper-
iments are: assessment of current focus functions ap-
plied on special biomedical data; a proposed modifi-
cation to existing autofocusing algorithm and its eval-
uation; and a method based on genetic programming
designed for finding a suitable focus function espe-
cially for similar image data.
1.1 Materials and Methods
The data used were TOPRO-dyed high-resolution im-
ages of desmocytes. They were acquired in either
confocal or standard (non-confocal) mode.
440
Luisa Brázdilová S. (2007).
TOWARDS RELIABLE AUTOFOCUSING IN AUTOMATED MICROSCOPY.
In Proceedings of the Fourth International Conference on Informatics in Control, Automation and Robotics, pages 440-447
DOI: 10.5220/0001639704400447
Copyright
c
SciTePress
Cut outs 200× 200 pixels
1
from 6 different sam-
ples were used for the experiments in order to reduce
the time consumption. Each sample was acquired 99
times, once for each z-position. The size of the step
between two successive images was manually cho-
sen so that all the information needed for experiments
would be captured. It was also dependent on whether
the confocal mode was used or not. The z-positions
are described by their natural order and treated as
nondimensional values without respect to the step size
used.
The information about the images is summarized
in Table 1. In the last column there are the z-positions
that are considered to be in focus. These numbers
come from human perception and will be used as a
reference.
Table 1: Information about image samples used for experi-
ments.
No. Step size (µm) Mode In-focus pos.
1 0.5 standard 53
2 0.5 confocal 33
3 0.2 confocal 43
4 0.2 confocal 42
5 0.5 confocal 47
6 1.0 standard 45
A technique called binning
2
(Netten, 1997;
Kozubek et al., 1999; Boddeke et al., 1994) was ap-
plied on the samples. It is useful not only for reducing
the spatial frequency, but it also minimizes the pro-
cessing time
3
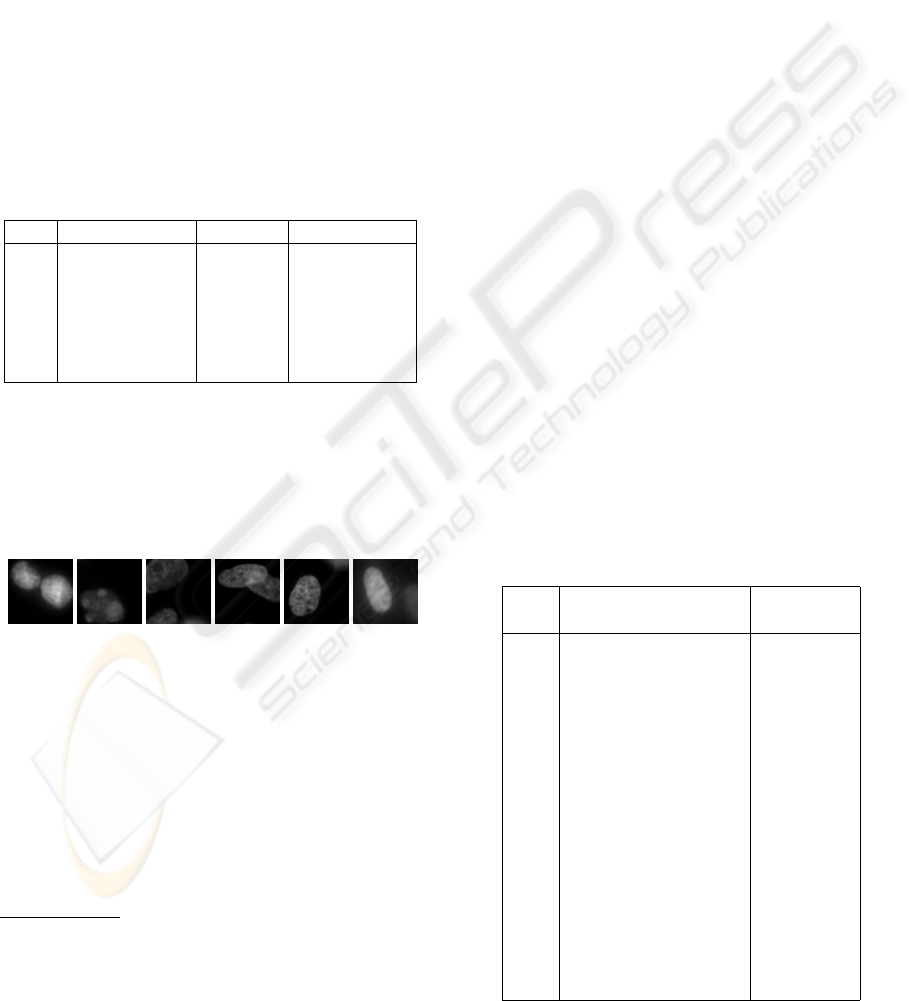
. The images after binning are presented
in Figure 1.
Figure 1: The in-focus images of testing samples 1 to 6 of
TOPRO-dyed high-resolution images of desmocytes, after
4× 4. binning.
2 STATE OF THE ART
In order to be able to implement an efficient and ac-
curate autofocusing, a combination of a good focus
function and a robust procedure for searching for the
maximum is needed.
1
This size is sufficient if suitable cut outs, i.e., those con-
taining cells or other objects of interest, are selected.
2
A group of adjacent pixes, here 4 × 4, is combined to
form a superpixel.
3
Now we have cut outs 50× 50 pixels.
2.1 Focus Functions
16 classic focus functions from the literature were
tested on the 6 samples (the list of these functions can
be found in the appendix). The functions were imple-
mented in MATLAB that is primarily optimized for
computations with matrices, and therefore they were
expressed using MATLAB-like functions with X be-
ing the acquired image. One of the main noticeable
changes is the introduction of 2D convolution instead
of pixel-wise operations that appear in the original no-
tation of the functions. More details about the new
notation will be given in section 4.
The error of a focus function is defined as a differ-
ence between the focus position given by this function
and the reference focus position provided by human
experts (see Table 1). By comparison of the errors of
the focus functions under test, and after visual evalua-
tion of the shape of these focus functions (with respect
to the desired properties) it was concluded that five of
them are applicable for such biomedical data. The
two most suitable focus functions are the Variance
(see Appendix for the formula) that was expressed in
the form
div(sum(pow(minus(X,avg(X)),2)),area(X))
and the Vollath’s F5 function (also in Appendix) ex-
pressed in the form
minus(sum(times(X,conv(X,V1))),
sum(times(X,conv(X,V2))))
where
V1=(1 0 0)
and
V2=(1 0 0 0 0)
.
Nevertheless, none of them is fully satisfactory.
The mean error is never less than 4, the total error on
the six samples is 24 or more. Detailed results can be
found in Table 2.
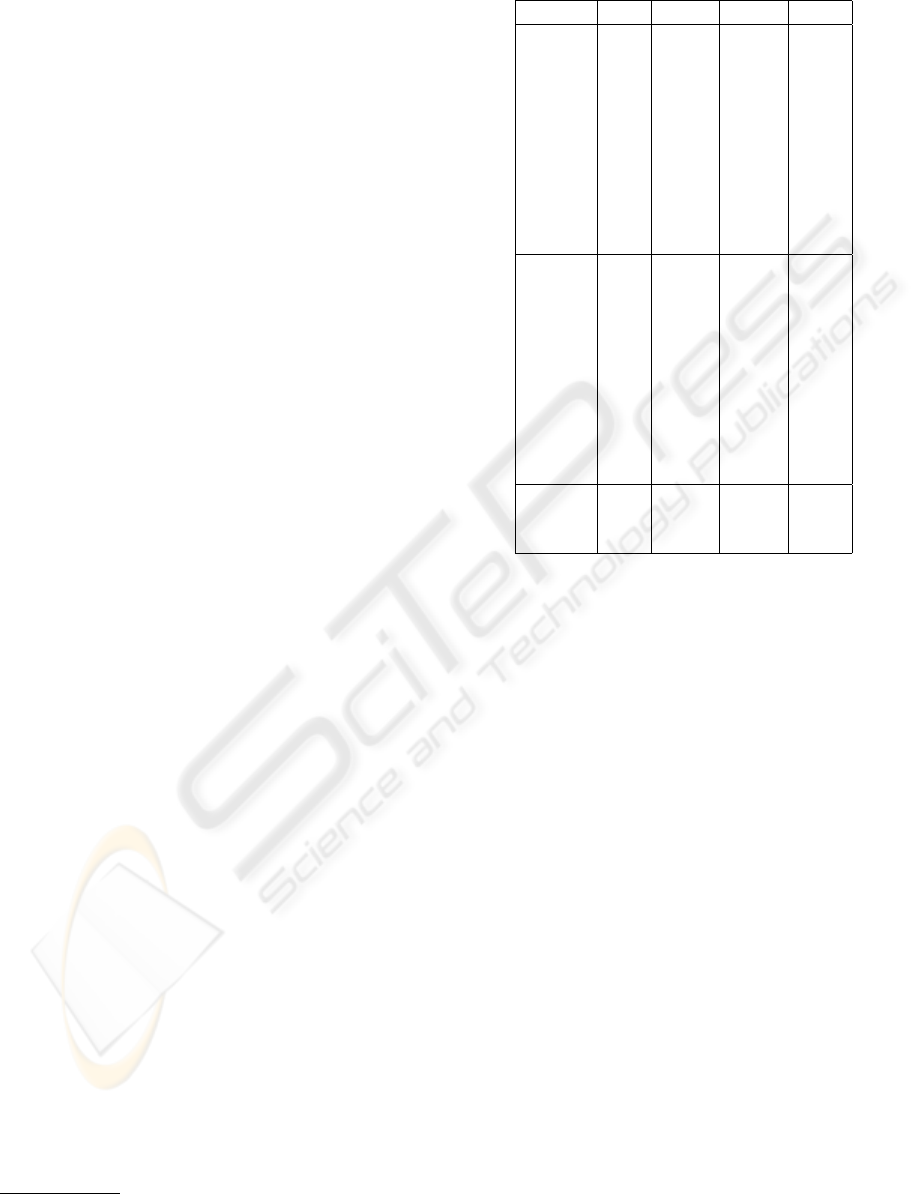
Table 2: Errors of the 16 focus functions.
Func. Samples Total Mean
No. 1 2 3 4 5 6 error error
1 15 3 2 0 1 11 32 5.3
2 12 2 22 1 2 25 64 10.6
3 12 2 2 0 2 6 24 4.0
4 44 3 0 1 2 23 73 12.2
5 46 3 2 1 1 25 78 13.0
6 12 2 2 1 2 23 43 7.2
7 26 3 0 0 2 17 48 8.0
8 15 2 2 0 2 7 28 4.6
9 49 1 21 41 46 33 191 31.8
10 15 2 2 0 2 7 28 4.6
11 49 1 0 41 46 33 170 28.6
12 49 66 18 3 45 33 247 41.2
13 49 17 18 1 46 34 165 27.5
14 12 2 2 0 2 6 24 4.0
15 15 2 2 0 2 7 28 4.6
16 24 48 41 57 48 53 271 45.2
TOWARDS RELIABLE AUTOFOCUSING IN AUTOMATED MICROSCOPY
441
2.2 Algorithm for Finding the
Maximum Focus
For the purpose of automatic focusing we tested Bod-
deke’s algorithm (Boddeke et al., 1994). It was
originally designed for unimodal focus functions and
works in three phases as follows:
The algorithm starts with the coarse phase at an
arbitrary (but known) position z
0
. First, two samples
are acquired and their focus values are compared in
order to determine the direction towards the focus.
This may be a problem for focus functions with lo-
cal minima, such as those applied on images with two
or more natural in-focus planes. Then, more images
are acquired until the last focus value is lower than
the previous one. All the samples of the first phase
are acquired evenly with the step ∆z
coarse
.
The starting position and the direction of the fol-
lowing fine phase, which is based on the same princi-
ple as the coarse phase, are given by the results of the
coarse phase. The size step used during the fine phase
is ∆z
fine
. Its size should correspond to the width of the
so-called quadratic region (see (Boddeke et al., 1994)
for details).
The last, refine phase of Boddeke’s algorithm con-
sists of capturing N
refine
images around the position
of the maximum focus function value found so far,
fitting a parabola and finding its maximum.
The tuning of such an algorithm is a difficult goal:
it concerns multiobjective optimization as we want to
find the values of several parameters (∆z
coarse
, ∆z
fine
,
N
refine
) while the number of images needs to be min-
imized and at the same time the accuracy and relia-
bility to be maximized. Moreover, the goal is to make
the algorithm robust to non-unimodal focus functions.
3 BODDEKE’S ALGORITHM
IMPROVEMENT
Here we present the proposed modification of the al-
gorithm described in 2.2:
1. In the coarse phase two images are acquired with a
step ∆z
coarse
in order to find the direction towards
the focus. Using the same step size, more images
are acquired until the last focus value is lower than
the previous one.
2. Interphase: the interval in which the focus lies is
assured by acquiring two images, one to each di-
rection, with a step ∆z
fine
from the z-position of
the highest focus so far. The position of the higher
focus function value of these two samples is the
direction towards the focus. This step may help
overcoming the bimodality that can occur.
3. In the fine phase images are acquired until the last
focus value is lower than the previous one, with
the step size ∆z
fine
from the position and to the
direction determined by the previous phase. The
two z-positions of the highest focus function val-
ues are passed to the refine phase.
4. The two z-positions from the fine phase together
with three more (one in between and two on the
sides of the interval, all being equidistant) are
used for parabola fitting. The maximum com-
puted from the parabola determines the position
of the maximum focus.
The following parameters were determined based on
empirical testing:
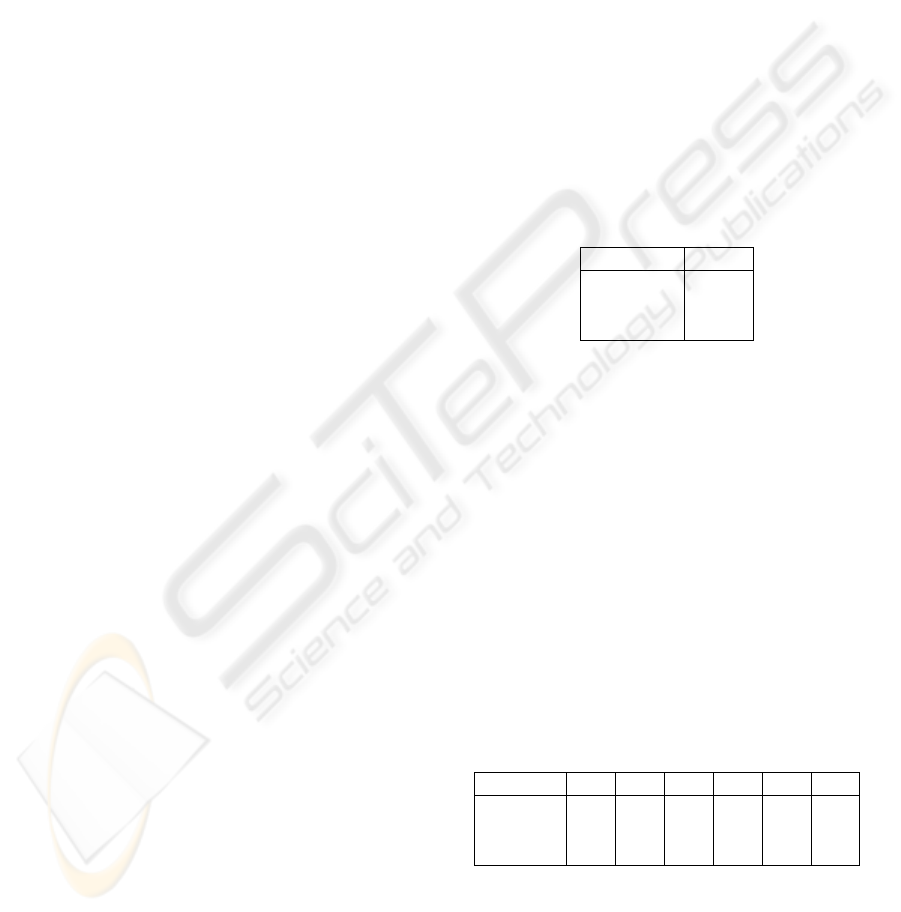
Table 3: Recommended parameters for the improved ver-
sion of Boddeke’s algorithm.
Parameter Value
∆z
coarse
10
∆z
fine
6
N
refine
5
The total number of images needed depends
strongly on the starting position z
0
. Yet some esti-
mation can be done, again based on some empirical
tests: it varies from 8 to 14. Naturally, starting near
to the in-focus position lowers the number of images
needed. The efficiency of this algorithm is shown
by the results of the algorithm applied together with
the Variance focus function on the all 6 samples pre-
sented in Table 4. For each sample, all 99 starting z-
positions were tested. The target is the position of the
actual maximum of the focus function, irrespectively
of the real in-focus position (we are assessing the al-
gorithm itself now, not the accuracy of the Variance
focus function, which is already known).
Table 4: Results of the improved version of Boddeke’s al-
gorithm using the Variance: ∆z
fine.
= 6, ∆z
coarse
= 10: a
comparison of target, mean and standard deviation values.
Sample 1 2 3 4 5 6
Target 68 35 41 42 49 52
Mean 68.9 35.1 42.2 41.9 48.2 53.5
Sted. dev. 0.30 0.54 0.60 0.30 1.18 4.62
It can be concluded that the improved Boddeke’s
algorithm is a reliable procedure for adaptive search-
ing the maximum of a function. If the focus function
was more accurate, we would have been able to locate
the position of the maximum focus quickly and pre-
cisely. Therefore, an effort should be made in order to
ICINCO 2007 - International Conference on Informatics in Control, Automation and Robotics
442
find better focus function since the focus function is
currently the weakest link of the whole autofocusing.
4 FINDING THE FOCUS
FUNCTION
The focus functions developed up to now are based
on previous knowledge about the differences in in-
formation content in focused and unfocused image.
But the space of all focus functions is potentially in-
finite. Nevertheless, we are looking for functions that
are easy to compute. It means that these functions
should be composed of relatively small number of op-
erators and operands, such as the focus functions that
were invented up to date. Still there may be plenty
of other similar functions with unthought-of perfor-
mance. Genetic programming (Koza, 1992) is a ro-
bust technique suitable for searching for optimal so-
lution in a large state space. We will use it in an at-
tempt to find a new focus function that will work well
on noisy biomedical images.
Genetic programming works in the same way as
genetic algorithms (Mitchell, 1996), but the popula-
tion is composed of functions (or programs), that can
be expressed in tree structure. The root represents the
output of the function, the leaves represent the argu-
ments. Inner nodes represent operators (here called
elementary functions).
Genetic programming, identically to the theory of
programming languages, faces the challenge of distin-
guishing between different data types. Strongly typed
genetic programming (Montana, 1995) is a branch of
genetic programming that finds inspiration in strongly
typed programming languages, when dealing with
this challenge. A modified version of this approach,
adapted especially for our data types needs, will be
presented here.
The MATLAB matrices of
doubles
are the only
data types used for our computations. However, we
need to work with matrices of any size, and distin-
guish between them. We will therefore define data
types by pairs of integers that will correspond to the
matrix sizes
4
. In addition, some functions have natu-
ral restrictions on the relations among the types (both
output and input). We will therefore define them as
generic functions (see (Montana, 1995)). Generic
functions are functions that are able to work with
more data types. They get instantiated during the tree
building. This approach needs special attention to be
paid during the process of tree generation (this will be
described in detail in next sections).
4
For example, a scalar will be of type (1, 1) etc.
Table 5: Table of functions, terminals and their types.
Function Arity Output Input 1 Input2
plus 2 a,b a,b a,b
minus 2 a,b a,b a,b
times 2 a,b a,b a,b
mtimes 2 a,c a,b b,c
nmtimes 2 a,b 1,1 a,b
mntimes 2 a,b a,b 1,1
divnum 2 a,b a,b 1,1
divmat 2 a,b a,b a,b
pownum 2 a,b a,b 1,1
conv 2 a,b a,b c,d
sum 1 1,1 a,b –
abs 1 a,b a,b –
avg 1 1,1 a,b –
log2 1 a,b a,b –
log10 1 a,b a,b –
uminus 1 a,b a,b –
area 1 1,1 a,b –
min 1 1,1 1,a –
max 1 1,1 1,a –
hist 1 256,1 a,b –
matrand 0 a,b – –
2 0 1,1 – –
X 0 x, y – –
If the type (or one of the pair components) is
generic, its value is represented by a symbolic value
denoted by alphabetic letters a, b, etc. The set of
symbolic values used for definition of a generic func-
tion represents the information we have at the moment
about relations among the type components. During
the process of creation of a new tree individual, there
is a moment when an elementary function is selected
for a node. If this elementary function is a generic
function, we may only define the input and output
types by symbolic values, but the important informa-
tion about the relation is preserved. For example, if a
binary function’s output must be of the same type as
its left-most input while the right-most input can be
arbitrary, we define output in terms of symbolic val-
ues (a, b), the left-most input also by (a, b), but the
right-most input is of type (c, d). The set of symbolic
values in this case is {a, b, c, d}. During instantiation,
each of these values is mapped to a specific numeric
value (e.g., 3, 8, 5 and 1), but the property that the
output type equals the right-most input’s type holds.
Of course, sets of symbolic values of different generic
elementary functions are totally independent (e.g., a
in the set of symbolic values of a function has noth-
ing to do with a belonging to the set of symbolic val-
ues of any other elementary function within the same
tree individual). Only when an elementary function
TOWARDS RELIABLE AUTOFOCUSING IN AUTOMATED MICROSCOPY
443

is assigned to a node that is direct descendant of an-
other one, its instantiated output type and the relevant
instantiated input type of the other function must fit.
The aim is to have all the nodes fully instantiated after
the tree is generated. The result of crossover, mutation
etc. must be a valid function too.
The list of functions and terminals used together
with their types specification is presented in Table 5.
The terminal set consists of a variable X that repre-
sents the image input (its size (x, y) is known in ad-
vance), and a constant 2 that occurs very often in the
classic focus functions. Function
matrand
of null par-
ity generates random matrices of desired size in the
program generation time. If the size is unspecified,
matrand
generates a matrix of random size between
(1, 1) and (10,10). Therefore its output type gets in-
stantiated on a random basis. Some functions have
predefined type values from the definition: for ex-
ample, the function
sum
produces the sum of the el-
ements of an arbitrary matrix and therefore its output
type is fixed and its value is (1, 1). The function
hist
computes the histogram with 256 bins. The function
area
gives the multiplication of matrix sizes,
conv
is
the 2D convolution of two matrices. The meaning of
other functions is straightforward.
4.1 Generation of a New Tree
The most complicated part of the process lies in man-
aging the data flow during the instantiation. The fol-
lowing definition will be necessary:
Definition 4.1 A generic function is type-consistent
if and only if it holds that if all the input types are
fully instantiated, the output type is fully determined
as well.
An example of a function that does not fulfil this con-
dition is a unary function of which input type is (a, b)
and the output type is (c, d). The algorithm proposed
can work only on type-consistent functions.
The tree is built recursively, in a depth-first man-
ner. The data structures used by the algorithm are:
The table of functions and terminals – The table
stores information about the generic functions and
their types in terms of symbolic or predefined val-
ues, such as shown in Table 5. It is only read dur-
ing the process.
The function instances in the nodes – This data
structure is created at the same time when a node
is created. The information gets updated gradu-
ally. First, the name of the assigned elementary
function and its arity is stored here. Every time
a type component of this function instance is
instantiated, the value is stored here as well.
Investigating the node, it can be assumed what
types have already been instantiated.
The information flow concerning the types is done in
four ways (Procedure A to D). Every time a new node
is created, its output type might, but does not have to,
be fully instantiated. (The root of any program that
aims to become a focus function falls among those
that have a specified output. It is the type (1, 1) be-
cause we want to measure a focus by a scalar value.)
Based on this information, a function is randomly se-
lected only from those that comply with the output
type request. The number of the direct descendants of
this node is now known.
The table of functions and terminals (Table 5) is
read first in order to get the predefined values of type
components, if there are any.
The components of the output type of the node are
checked. If any of them is already instantiated, the
symbolic values in the table are checked. If there is a
symbolic value for this component of the output type,
it means that we have instantiated it already. The spe-
cific value gets in the node to every type component
that has the same symbolic value in the table (proce-
dure B). The process can cause (partial) instantiation
of both the input and output types of the node.
The information known at that moment is pro-
cessed further when the left-most descendant is gen-
erated.
When the left-most subtree is completed, the pro-
cess of elaborating this node continues. Because of
the type consistence property the output type of the
left descendant must be fully instantiated at that mo-
ment. If the left input type has not been instanti-
ated before, it takes the value of the left-most descen-
dant’s output (Procedure C). This information may
enrich the information about other types in the ac-
tual node, therefore procedure D is needed: it goes
through the input types and if some of their values are
newly instantiated, the symbolic values in the table
are checked similarly to procedure B. The informa-
tion goes to other components (including the output
types) according to the symbolic value in the table.
The same process continues with all other descen-
dants, including procedures C and D. At the latest
when the last branch is finished, the output type of
the actual node is fully instantiated, owing to the type
consistent property and procedure D.
4.2 Genetic Programming Experiment
Description
We used GPLAB (Silva and Almeida, 2003b), a ge-
netic programmming toolbox for MATLAB, which
ICINCO 2007 - International Conference on Informatics in Control, Automation and Robotics
444

we modified in order to be able to handle strong typ-
ing. The initial population was generated using the
ramped half-and-half method (Koza, 1992). Dynamic
maximum tree depth, a technique for avoiding bloat
(Silva and Almeida, 2003a), was also incorporated.
The population size was 20, the dynamic limit and
maximum limit were 5 and 20, respectively.
Standard genetic operators
reproduction
,
mutation
and
crossover
were used. The type
constraint was resolved easily for mutation: a node
was randomly selected, its type was found and a
random tree was generated whose root’s output type
was the type needed. Crossover was performed only
if the node selected from the first parent was present
at least once in the second parent as well. In the case
of multiple occurrence the final node for exchange
was selected randomly.
The lexicographic parsimony pressure was used
for selection together with elitism.
Fitness function was computed in the following
way: first, candidate focus function was evaluated
on the whole training set (an image sample acquired
with various z-positions). Then a maximum was
found, and the z-position that exhibits the maximum
value was compared with the reference z-position.
The difference is the error and therefore should be
minimized. To eliminate functions that are constant
around their maxima, the number of z-positions ex-
hibiting the maximum value was added to the fitness
function to disrate functions with undesirable shape.
More training sets are needed in order to prevent
the genetic algorithm from guessing the correct z-
position independently on the image itself. Due to
high time consumption, the training set of three sam-
ples was used. It concerned samples 1, 3 and 6. The
remaining three samples were used for testing.
4.3 Results
The resulting functions were then compared with one
of the best classic functions, i. e. Variance, on a test-
ing set of three different samples. Five new functions
were giving comparable results. They even outper-
formed them on some samples. Their accuracy, i. e.,
how far a maximum was identified from the real one
on the z axis, was evaluated together with their shapes
and brevity.
One of the most interesting results is:
sum(plus(times(times(X,X),X),X))
which can be mathematically expressed as
f(z) =
∑
x
∑
y
((I(x, y,z))
3
+ I(x, y, z))
The focus position given by that function is compared
to the output of Variance in Table 6. In three cases
(samples 3, 4 and 5) the genetic programming func-
tion was as successful as the two classic functions.
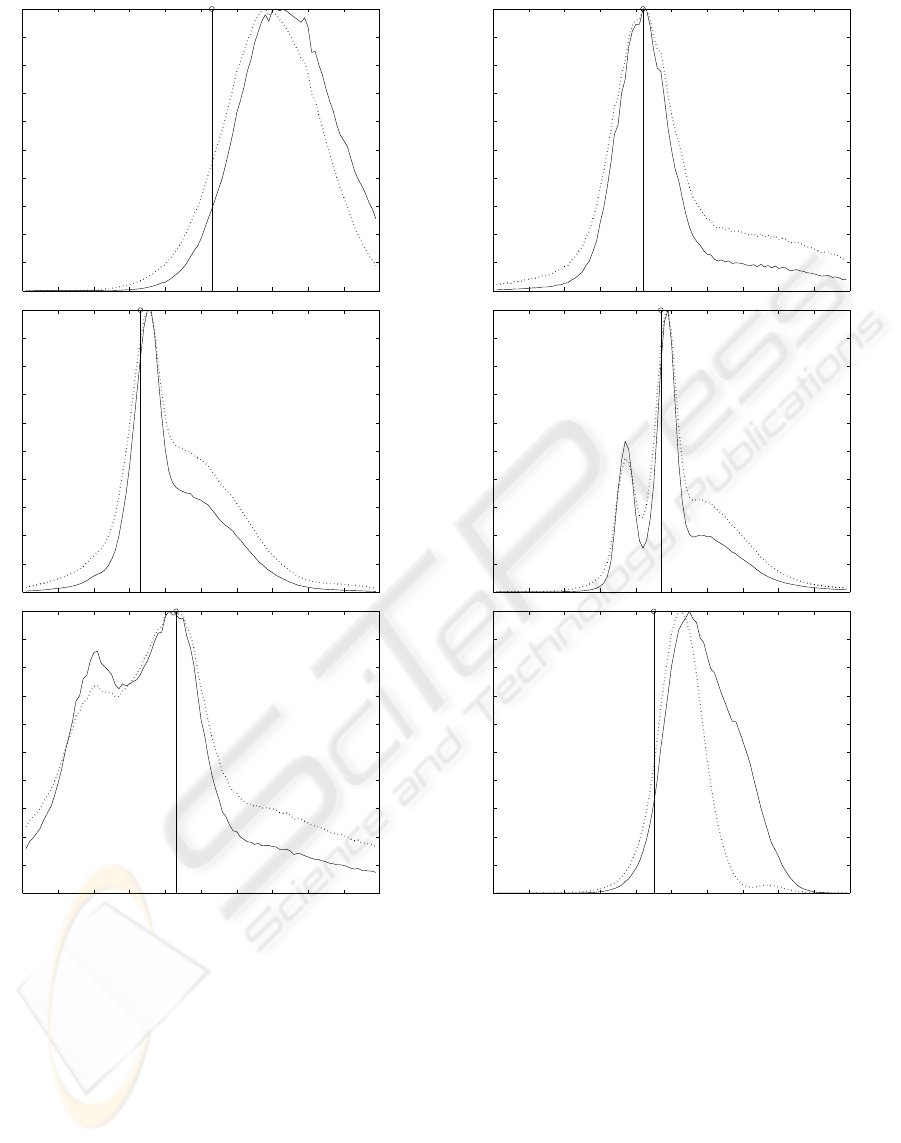
The graphs of the behaviour of this new focus
function on all six samples can be found in Figures
2 and 3.
Table 6: Comparison of focus positions found by Variance
and the genetic result.
Sample Reference Variance Genetic result
1 53 68 71
2 33 35 36
3 43 41 41
4 42 42 42
5 47 49 49
6 45 52 55
Total error – 28 35
Mean error – 4.6 5.8
5 CONCLUSION
In this work several different tasks were accom-
plished. Their unifying topic is the autofocusing for
automated microscopy that was simulated on a per-
sonal computer. First, 16 classic functions were tested
on biomedical data. None of them was as accurate and
reliable as would be necessary. Nevertheless, some
of them are usable, mainly Variance and Vollath’s F5.
Second, Boddeke’s algorithm was tested. It turned out
that the algorithm can be modified so that it reliably
finds the maximum of the function despite the poten-
tial bimodality. Some 10 acquired images are needed
in average.
The weakest link is certainly the focus function it-
self. If this issue is solved, the automated focusing
can become routine work without the need of human
assistance. In order to find such a function, a pre-
liminary genetic programming experiment was con-
ducted. Its result is comparable to the Variance. This
genetic programming application has shown the im-
portance of solving the strong typing issue. An algo-
rithm for tree generation using a variant of strongly
typed genetic programming was designed, and an im-
portant property – the type-consistence – of elemen-
tary functions was recognized and defined.
5.1 Future Work
The genetic programming design could be improved
by more careful selection of elementary functions, pa-
rameters etc. Running the program parallelly could
enable using more individuals, generations and train-
ing samples. This altogether could result in a highly
TOWARDS RELIABLE AUTOFOCUSING IN AUTOMATED MICROSCOPY
445
0 10 20 30 40 50 60 70 80 90 100
0
0.1
0.2
0.3
0.4
0.5
0.6
0.7
0.8
0.9
1
0 10 20 30 40 50 60 70 80 90 100
0
0.1
0.2
0.3
0.4
0.5
0.6
0.7
0.8
0.9
1
0 10 20 30 40 50 60 70 80 90 100
0
0.1
0.2
0.3
0.4
0.5
0.6
0.7
0.8
0.9
1
Figure 2: Genetic result (solid line) and Variance (dotted
line) on testing samples 1 to 3 from top to bottom. The
reference value is highlighted.
fit focus function of unprecedented performance that
would discover the hidden features of images in fo-
cus. Finally, the results from PC simulations could be
realized on a real automated microscope.
ACKNOWLEDGEMENTS
This work was supported by the Ministry of Edu-
cation of the Czech Republic (Grants No. MSM-
0 10 20 30 40 50 60 70 80 90 100
0
0.1
0.2
0.3
0.4
0.5
0.6
0.7
0.8
0.9
1
0 10 20 30 40 50 60 70 80 90 100
0
0.1
0.2
0.3
0.4
0.5
0.6
0.7
0.8
0.9
1
0 10 20 30 40 50 60 70 80 90 100
0
0.1
0.2
0.3
0.4
0.5
0.6
0.7
0.8
0.9
1
Figure 3: Genetic result (solid line) and Variance (dotted
line) on testing samples 4 to 6 from top to bottom. The
reference value is highlighted.
0021622419, No. LC535 and No. 2B06052).
REFERENCES
Boddeke, F. R. (1999). Quantitative Fluorescence Mi-
croscopy. PhD thesis, Technische Universiteit Delft.
Boddeke, F. R., van Vliet, L. J., Netten, H., and Young, I. T.
(1994). Autofocusing in microscopy based on the otf
and sampling. Bioimaging, 2:193–203.
ICINCO 2007 - International Conference on Informatics in Control, Automation and Robotics
446

Bueno-Ibarra, M. A.,
´
Alvarez Borrego, J., Acho, L., and
Ch
´
avez-S
´
anchez, M. C. (2005). Fast autofocus algo-
rithm for automated microscopes. Optical Engineer-
ing, 44.
Firestone, L., Cook, K., Culp, K., Talsania, N., and Preston,
K. (1991). Comparison of autofocus methods for au-
tomated microscopy. Cytometry, 12:195–206.
Geusebroek, J., Cornelissen, F., Smeulders, A., and Geerts,
H. (2000). Robust autofocusing in microscopy. Cy-
tometry, 39:1–9.
Groen, F. C., Young, I. T., and Ligthart, G. (1985). A com-
parison of different focus functions for use in autofo-
cus algorithms. Cytometry, 6:81–91.
Koza, J. R. (1992). Genetic Programming: On the Pro-
gramming of Computers by Means of Natural Selec-
tion. MIT Press.
Kozubek, M., Kozubek, S., Luk
´
a
ˇ
sov
´
a, E., Mare
ˇ
ckov
´
a, A.,
B
´
artov
´
a, E., Skaln
´
ıkov
´
a, M., and Jergov
´
a, A. (1999).
High-resolution cytometry of fish dots in interphase
cell nuclei. Cytometry, 36:279–293.
Mitchell, M. (1996). An Introduction to Genetic Algo-
rithms. Massachusetts Institute of Technology.
Montana, D. J. (1995). Strongly typed genetic program-
ming. Evolutionary Computation, 3:199–230.
Netten, H. (1997). Automated Image Analysis of FISH-
Stained Cell Nuclei. PhD thesis, Delft University of
Technology.
Santos, A., Ortiz de Sol
´
orzano, C., Vaquero, J. J., Pe
˜
na,
J. M., Malpica, N., and del Pozo, F. (1997). Evalu-
ation of autofocus functions in molecular cytogenetic
analysis. Journal of Microscopy, 188:264–272.
Silva, S. and Almeida, J. (2003a). Dynamic maximum tree
depth - a simple technique for avoiding bloat in tree-
based gp. In Proceedings of the Genetic and Evolu-
tionary Computation Conference (GECCO-2003).
Silva, S. and Almeida, J. (2003b). Gplab - a genetic pro-
gramming toolbox for matlab. In Proceedings of
the Nordic MATLAB Conference (NMC-2003), pages
273–278.
Sun, Y., Duthaler, S., and Nelson, B. J. (2004). Autofocus-
ing in computer microscopy: Selecting the optimal fo-
cus algorithm. Microscopy Research and Technique,
65:139–149.
APPENDIX
I(x, y, z) ...The intensity of a pixel at a position (x, y)
in an image I acquired at the z-position z
m, n .. .The width and height of an image I
I(z) ...The average of the pixel intensity of an image
I acquired at the z-position z, i. e.,
∑
x
∑
y
I(x,y,z)
m·n
h(i) ...Number of pixels of intensity equal to i in I
p(i) ... The frequency of occurrence of pixels with
intensity equal to i in an image I, i. e., p(i) =
h(i)
mn
The list of the focus functions tested:
1. Absolute Gradient
f(z) =
∑
x
∑
y
|I(x, y, z) − I(x, y− 1, z)|
2. Square gradient
f(z) =
∑
x
∑
y
(I(x, y, z) − I(x, y− 1, z))
2
3. Netten’s filter
f(z) =
∑
x
∑
y
(I(x+ 1, y, z) − I(x− 1, y, z))
2
4. Energy Laplace
f(z) =
∑
x
∑
y
C(x, y,z)
2
where C(z) = I(z) ∗
−1 −4 −1
−4 20 −4
−1 −4 −1
5. Laplacian
f(z) =
∑
x
∑
y
(I(x, y− 1, z) − 2I(x, y, z) + I(x, y+ 1, z))
2
6. Tenegrad’s function
f(z) =
∑
x
∑
y
S
x
(x, y, z)
2
+ S
y
(x, y, z)
2
where
S
x
(z) = I(z)∗
1 2 1
0 0 0
−1 −2 −1
, S
y
(z) = I(z)∗
−1 0 1
−2 0 2
−1 0 1
7. Signal Power
f(z) =
∑
x
∑
y
(I(x, y, z)
2
8. Variance
f(z) =
1
mn
∑
x
∑
y
(I(x, y, z) −
I(z))
2
9. Normalized Variance
f(z) =
1
mn(I(z))
2
∑
x
∑
y
(I(x, y, z) −
I(z))
2
10. Absolute Variance
f(z) =
1
mn
∑
x
∑
y
|(I(x, y,z) −
I(z)|)
11. Normalized Absolute Variance
f(z) =
1
mn(I(z))
2
∑
x
∑
y
|(I(x, y,z) − I(z)|)
12. Histogram Range
f(z) = max
i
(h(i) > 0) − min
i
(h(i) > 0)
13. Histogram Entropy
f(z) = −
∑
i
p
i
· log
2
p
i
14. Vollath’s F
4
f(z) =
∑
x
∑
y
I(x, y, z) · I(x+ 1, y, z)−
−
∑
x
∑
y
I(x, y, z) · I(x+ 2, y, z)
15. Vollath’s F
5
f(z) =
∑
x
∑
y
I(x, y, z) · I(x+ 1, y, z) − mn(
I(z))
2
16. Spectral Analysis
f(z) =
∑
i
p
i
· log
10
i
TOWARDS RELIABLE AUTOFOCUSING IN AUTOMATED MICROSCOPY
447
