
Med2Meta: Learning Representations of Medical Concepts with
Meta-embeddings
Shaika Chowdhury
1
, Chenwei Zhang
2
, Philip S. Yu
1
and Yuan Luo
3
1
Department of Computer Science, University of Illinois at Chicago, Chicago, Illinois, U.S.A.
2
Amazon, Seattle, Washington, U.S.A.
3
Department of Preventive Medicine, Northwestern University, Chicago, Illinois, U.S.A.
Keywords:
Representation Learning, Electronic Health Records, Meta-embeddings, Graph Neural Networks.
Abstract:
Distributed representations of medical concepts have been used to support downstream clinical tasks recently.
Electronic Health Records (EHR) capture different aspects of patients’ hospital encounters and serve as a rich
source for augmenting clinical decision making by learning robust medical concept embeddings. However,
the same medical concept can be recorded in different modalities (e.g., clinical notes, lab results) — with each
capturing salient information unique to that modality — and a holistic representation calls for relevant feature
ensemble from all information sources. We hypothesize that representations learned from heterogeneous data
types would lead to performance enhancement on various clinical informatics and predictive modeling tasks.
To this end, our proposed approach makes use of meta-embeddings, embeddings aggregated from learned
embeddings. Firstly, modality-specific embeddings for each medical concept is learned with graph auto-
encoders. The ensemble of all the embeddings is then modeled as a meta-embedding learning problem to
incorporate their correlating and complementary information through a joint reconstruction. Empirical results
of our model on both quantitative and qualitative clinical evaluations have shown improvements over state-of-
the-art embedding models, thus validating our hypothesis.
1 INTRODUCTION
With the increase in healthcare efficiency and im-
provement in patient care that comes with archiving
medical information digitally, more healthcare facili-
ties have started adopting Electronic Health Records
(EHRs) (Shickel et al., 2018; Knake et al., 2016;
Charles et al., 2013). EHRs store a wide range of
heterogeneous patient data (e.g., demographic infor-
mation, unstructured clinical notes, numeric labora-
tory results, structured codes for diagnosis, medica-
tions and procedures) as a summary of the patient’s
entire hospital stay. As a result of the richness of med-
ical content that can be mined from EHR, its use in
predictive modeling tasks in the medical domain has
become ubiquitous.
How good a machine learning algorithm performs
is dependant on the representations of the data that
it is able to learn (Bengio et al., 2013). However, the
high dimensionality, diversity of data and complex as-
sociations between clinical variables in EHR means
the intermediate data representation learning task is
not trivial. Feature engineering with the help of a do-
main expert was prevalent in earlier works to specify
which clinical variables from the EHR to consider as
the input features (Jensen et al., 2012). Although the
features could be more precise as hand-picked with
domain knowledge, the manual effort, scalability is-
sues and inability to generalize render such approach
undesirable (Miotto et al., 2016).
Distributed representations of words, known as
word embeddings, have brought immense success in
numerous Natural Language Processing (NLP) tasks
(Goldberg, 2016). Recent works in deep learning ap-
plications for EHR (Choi et al., 2016c; Choi et al.,
2016b; Tran et al., 2015; Choi et al., 2016a; Choi
et al., 2017) emulated the concept of word embed-
ding as learning vector representations of medical
concepts. They focused on learning the embeddings
from predominantly one modality (e.g., unstructured
notes or structured clinical events) that excludes rel-
evant information about medical concepts found in
other modalities. For example, a patient’s symptoms
for a disease can be mentioned in physician notes but
could be missing from their structured clinical event
data of clinical codes. However, trying to fuse infor-
Chowdhury, S., Zhang, C., Yu, P. and Luo, Y.
Med2Meta: Learning Representations of Medical Concepts with Meta-embeddings.
DOI: 10.5220/0008934403690376
In Proceedings of the 13th International Joint Conference on Biomedical Engineering Systems and Technologies (BIOSTEC 2020) - Volume 5: HEALTHINF, pages 369-376
ISBN: 978-989-758-398-8; ISSN: 2184-4305
Copyright
c
2022 by SCITEPRESS – Science and Technology Publications, Lda. All rights reserved
369

mation from different modalities in EHR presents the
following obstacles for representation learning,
1. Inconsistency in Medical Concept Terminol-
ogy. In structured clinical events, the medical con-
cept is represented with ICD-9/ICD-10 clinical code.
While in unstructured clinical notes, it is either men-
tioned with a formal medical term for the concept or
an informal analogous term/phrase implicitly. It is
difficult to consistently detect the presence of a medi-
cal concept across different modalities.
2. Varying Contexts. Prediction based embed-
ding approaches (Choi et al., 2016b; Choi et al.,
2016a) work well on structured modality where each
patient can be represented as a sequence of visits of
codes, and can consider context in terms of the other
neighboring codes within the same visit. However, for
unstructured clinical notes, the context can be noisy
due to the presence of text describing all aspects of a
patient’s admission (e.g., past medical history).
3. Feature Associations Complexity. Other
types of patient information such as demographics
and laboratory results are possibly important signals
in prediction tasks. However, modeling their non-
linear and complex relations with the medical con-
cepts is not straightforward.
4. Interpretability. The resulting learned em-
bedding should be understandable as clinicians want
to know the underlying reasons for predictive results,
that should also comply with medical knowledge.
To address these obstacles our proposed approach,
Med2Meta, is formulated as a meta-embedding learn-
ing problem. Meta-embeddings are embeddings ob-
tained from the ensemble of existing embeddings. As
embeddings vary in terms of the corpus they are cre-
ated from and the approach used to learn them, (Chen
et al., 2013; Yin and Sch
¨
utze, 2015) have found that
they capture different semantic characteristics. Meta-
embedding learning exploits this idea and combines
the semantic strengths of different types of embed-
dings, which has been shown to outperform single
embedding on tasks such as word similarity and anal-
ogy (Yin and Sch
¨
utze, 2015). Med2Meta differs from
traditional meta-embedding learning in that it does
not combine existing pre-trained embeddings to ob-
tain the new embedding for each medical concept, as
most of these models are designed for one data modal-
ity. Rather, feature-specific embedding for each med-
ical concept in the EHR is first learned with graph
auto-encoder by considering each heterogeneous data
type as a different view. Using graph auto-encoder
for learning embeddings gives us the benefit of being
able to model the relations between different types of
medical concepts through the graph’s structure and,
at the same time, infuse relevant feature information
for each medical concept collected from a particu-
lar modality. In particular, with each medical con-
cept modeled as a node in the graph and edges be-
tween two nodes signifying corresponding relation-
ship found in EHR data w.r.t a modality, embeddings
are learned by considering features extracted from
that modality as a separate view. Med2Meta consid-
ers three different heterogeneous data types in EHR
as separate views — demographic information (dem),
laboratory results (lab) and clinical notes (notes).
Each graph is constructed with a novel graph con-
struction approach, where unique medical concepts
from structured clinical events are extracted to serve
as the nodes of the graph.
The embeddings learned from each view are then
considered as input sources for the meta-embedding
process, which are fused together using Dual Meta-
Embedding Autoencoders (Dual-MEAE). The en-
coders in Dual-MEAE are inspired by the idea in
(Roweis and Saul, 2000), where each data point and
its neighbors can be expected to lie on or close to a
locally linear patch of the manifold, where similar in-
stances should result in similar positions in the em-
bedding space. Therefore in Dual-MEAE, a pair of
encoders are allocated for each view — one for the
source embedding and the other for the average em-
bedding of the most similar medical concepts based
on that view —- in order to project different modal-
ities to a common meta-embedding space. That is,
the embedding of a medical concept can be expected
to be semantically similar to that of its most similar
medical concepts and, hence, should also lie nearby
in the meta-embedding space. To enable this, a sin-
gle decoder jointly tries to reconstruct the original in-
put source and the average embedding from the latent
meta-embedding representation. By minimizing the
reconstruction errors across all the views in an unified
manner, the intermediate meta-embedding represen-
tation is able to retain correlating information among
medical concepts within the same view, as well as
capture complementary information across different
views, hence learning a holistic vector representation
for each medical concept. In general, Med2Meta’s
contributions are threefold,
• Learns holistic embeddings of medical concepts
that fuses information from heterogeneous data types
in EHR through a meta-embedding process.
• Modality-specific embeddings capture the se-
mantic features of each EHR data type and are gen-
erated using Graph Auto-Encoders.
• Outperforms current state-of-the-art models in
both qualitative and quantitative experiments for a
publicly available EHR dataset.
HEALTHINF 2020 - 13th International Conference on Health Informatics
370
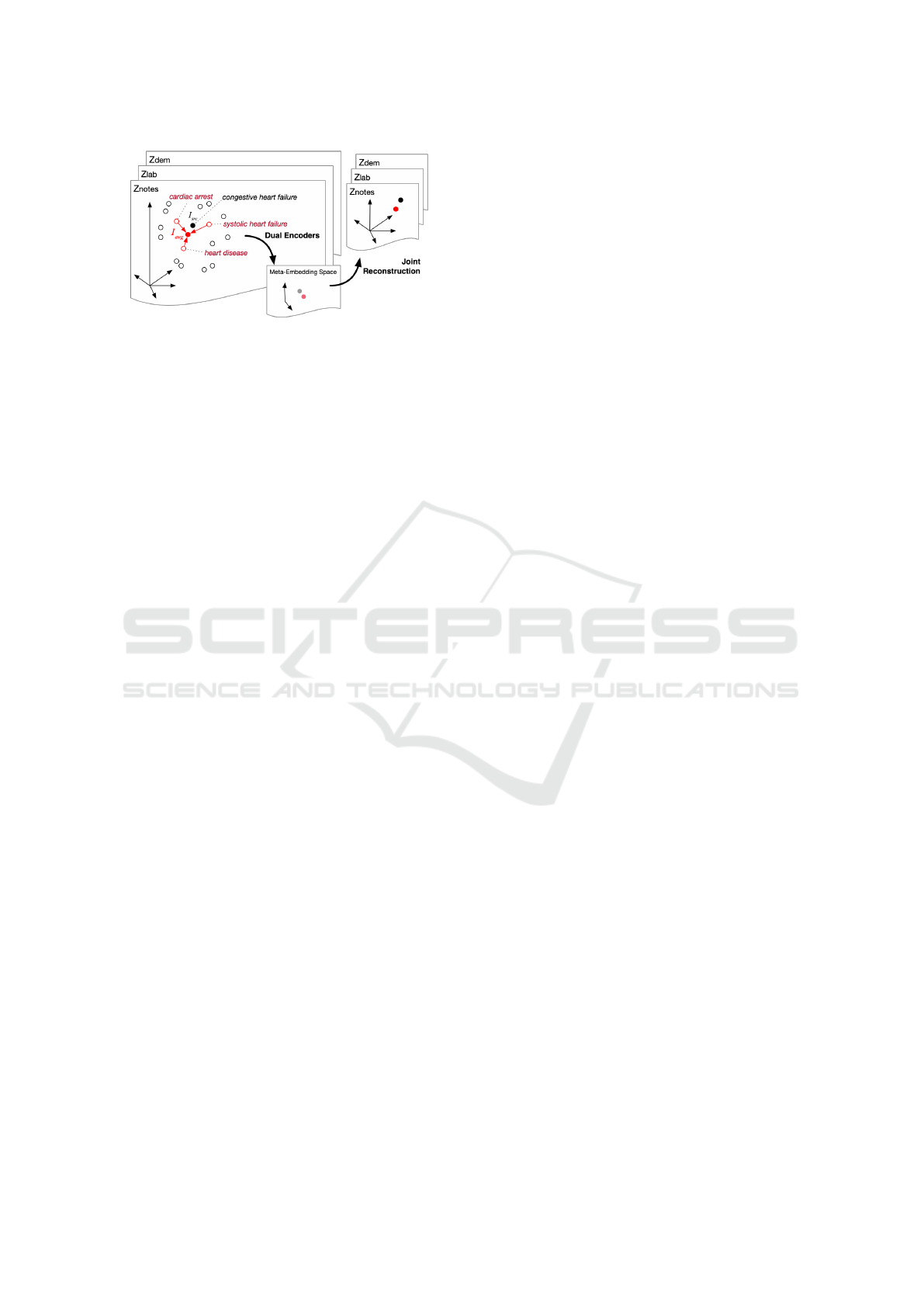
Figure 1: Proposed Dual-MEAE Model: Fuses embeddings
from different views with dual encoders for each view and
a single decoder to reconstruct jointly across all views.
2 META-EMBEDDING PROBLEM
DEFINITION
Let S
i
∈ R
V
i
×d
i
represent the i-th source medical con-
cept embedding, where i ∈ {1, ... k}, for total k = 3
source embeddings representing the three different
views in our case. Here V
i
is the medical concept
vocabulary covered by the i-th source embedding. In
our case, V
i
for all three sources i ∈ {1, .. . k} are the
same as the total number of unique medical concepts,
V, used to create each modality-specific graph is the
same, and hence we consider V
i
= V. The dimen-
sion of medical concept embedding for the respective
source is denoted by d
i
, where we set all d
i
to be of
the same dimension, d. The meta-embedding learn-
ing problem is then defined as learning an embedding
m(c) ∈ R
b
of dimension b in the meta-embedding
space for each medical concept c ∈ V .
3 Med2Meta ARCHITECTURE
Our proposed approach comprises two steps: 1) ob-
taining modality-specific embeddings and 2) meta-
embedding learning. In step 1, Graph Auto-Encoder
is trained on each modality features in turn to gen-
erate feature-specific embeddings to be considered as
input sources into step 2. While in step 2, all the in-
put source embeddings are ensembled through meta-
embedding learning to obtain the new embedding for
each medical concept.
3.1 Generating Modality-specific
Embeddings
The embedding learned for each medical concept
should holistically capture relevant features from each
data modality in EHR, as patient information cover-
age is not the same across all the modalities. For in-
stance, information regarding symptoms experienced
during a disease, its severity and other important ob-
servations made by physicians/ nurses are probably
only found in clinical notes. Hence, learning the
embeddings only from structured clinical records is
a sub-optimal approach. We use the same Graph
Auto-Encoder technique as used in (Chowdhury et al.,
2019), however, in this work graph for each view is
constructed using EHR information collected from all
the patients.
3.2 Fusion through Meta-embedding
Learning
The three types of modality-specific embeddings ob-
tained through graph auto-encoder exist in different
vector spaces and capture semantic relations of the
respective view. In order to learn an embedding
that collectively reflects semantic associations across
different modalities in EHR for each medical con-
cept, semantic knowledge from all the feature-specific
embeddings need to be integrated. Performing the
fusion by casting it as a meta-embedding problem
through autoencoding is meaningful as we intend to
learn the ensembled embedding as a meta-embedding
by reconstructing the modality-specific embeddings
jointly. The projection to a common meta-embedding
space would therefore lead to coherent enforcement
of mutual and correlative information present in the
embeddings. The proposed Dual-MEAE model is de-
picted in Figure 1.
We consider each modality specific embedding,
extracted from a view in step 1, as an input source
embedding, S
i
, into Dual-MEAE. In each source em-
bedding space, medical concepts that are semantically
similar in that view would exist geometrically closer
and thus can be expected to have similar distributed
vectors. So, our proposed approach reconstructs each
medical concept jointly from both the source embed-
ding and average embedding of its most similar med-
ical concepts.
Each component of our meta-embedding model,
Dual-MEAE, is discussed in detail in the following
sub-sections.
3.2.1 Encoder
On the encoder side, a set of dual encoders, Enc
i
src
and Enc
i
avg
, is set aside for each view i from i ∈
{dem, lab, notes}. For each medical concept c, its
learned source embedding with respect to view i and
the average of the source embeddings of most simi-
lar medical concepts to c in that view are fed as in-
Med2Meta: Learning Representations of Medical Concepts with Meta-embeddings
371

puts, I
i
src
(c) ∈ R
d
and I
i
avg
(c) ∈ R
d
, into Enc
i
src
and
Enc
i
avg
respectively. That is, the output feature ma-
trix Z
i
∈ R
V ×d
obtained for each view from step 1
is considered as the respective source embedding, S
i
,
for that view. Henceforth, Z
i
[c,:], which is the c-th
row of Z
i
for c ∈ V and represents the GAE-generated
distributed embedding vector of c-th medical concept,
is fed as input I
i
src
(c) into Enc
i
src
for the respective
view. From our preliminary results, we found that
projecting the output feature matrix from dimension
N × d → V × d
0
where d
0
< d through principal com-
ponent analysis (PCA) (Wold et al., 1987) improves
performance; so we feed embedding vector extracted
from the PCA output matrix Z
i
0
instead.
On the other hand, the accumulation of the source
embeddings of the most similar medical concepts to
c with respect to view i is computed as their aver-
age, avg
i
(c), and is considered as the input, I
i
avg
(c),
to the second encoder, Enc
i
avg
. We specifically chose
to compute the average of the neighborhood embed-
dings as it has shown to perform well in recent stud-
ies (Coates and Bollegala, 2018). To find the most
similar medical concepts of concept c, the top three
medical concepts sorted by highest values of dice co-
efficient for that view computed in step 1 are selected.
Top three gave us best results based on hyperparame-
ter analysis on validation set.
The dual encoders encode their inputs I
i
src
and I
i
avg
,
so that latent attributes and their non-linear relation-
ships could be effectively learned. Let us call the en-
coded representations from the dual encoders for each
medical concept as their target embeddings, T
i
src
∈ R
d
0
and T
i
avg
∈ R
d
0
, defined as
T
i
src
(c) = Enc
i
src
(I
i
src
(c)) (1)
T
i
avg
(c) = Enc
i
avg
(I
i
avg
(c)) (2)
Each encoder is implemented as a fully connected
neural network layer, with d
0
hidden units and ReLU
(Nair and Hinton, 2010) activation.
3.2.2 Meta-embedding Space
We first concatenate the respective encoded input rep-
resentations as T
i
(c) ∈ R
2d
0
,
T
i
(c) = [T
i
src
(c); T
i
avg
(c)] (3)
here [] denotes concatenation.
To get meta-embedding m(c) ∈ R
b
, each T
i
(c) is
first passed through one dense layer to model it to
respective intermediate representation, m
i
(c) ∈ R
2d
0
.
The concatenation of all m
i
(c) is considered as the
meta-embedding m(c), defined as,
m(c) = [m
i
(c)] (4)
for i ∈ {dem,lab,notes} and dimension b = 6d
0
.
As Dual-MEAE jointly encodes and decodes
across all the views, the meta-embedding is updated
with view-specific feature information in a meta-
embedding space. Thus, the meta-embedding space
will eventually be able to capture a holistic semantics
comprising of associations between medical concepts
within same as well as different views.
3.2.3 Decoder
In order to decode the semantically composite meta-
embedding representation, a single decoder Dec
maps the latent representation, m(c), from the meta-
embedding space back to corresponding source and
average embeddings across all views simultaneously.
The decoder is implemented as a neural network of
dense layers, each with d
0
hidden units and ReLu ac-
tivations. The output of the decoder is defined as,
˙
Dec(c) = Dec(m(c)). (5)
˙
Dec(c) ∈ R
6×d
0
contains the reconstructed source
and average embeddings for the three views.
This joint decoding from the meta-representation
simultaneously across all the views drives Med2Meta
in embedding both mutually related and complemen-
tary information of heterogeneous data types into the
meta-embedding space.
3.2.4 Objective Function
The meta-representation, m(c), optimized through the
minimization of the reconstruction losses across all
the views is taken as the final embedding for the cor-
responding medical concept. The overall objective
function for total k views and V medical concepts in
the training set is defined as,
L =
∑
c∈V
∑
i∈k
(ω
1
T
i
src
(c) − T
i
avg
(c)
2
+
ω
2
˙
Dec(c)[i
src
,:] − T
i
src
(c)
2
+
ω
3
˙
Dec(c)[i
avg
,:] − T
i
avg
(c)
2
)
(6)
˙
Dec(c)[i
avg
,:] and
˙
Dec(c)[i
src
,:] refer to the de-
coded embeddings for the average and source embed-
dings respectively of view i for medical concept c.
The first term in the loss is responsible for infusing
common associations between a medical concept and
other similar medical concepts, whereas, the second
and third try to preserve information essential locally
with respect to each view for each medical concept
HEALTHINF 2020 - 13th International Conference on Health Informatics
372

Table 1: Data Statistics Summary of MIMIC-III.
MIMIC-III
# of patients 7,499
# of visits 19,911
avg. # of visits per patient 2.66
# of unique ICD9 codes 4,893
avg. # of codes per visit 13.1
max # of codes per visit 39
during the respective reconstructions. Moreover, the
degree of these properties in the loss can be controlled
through the values of the coefficients ω.
4 EXPERIMENTAL SETUP
4.1 Source of Data
We evaluate our model on the publicly available
MIMIC-III dataset (Johnson et al., 2016). This
database contains de-identified clinical records for
> 40K patients admitted to critical care units over
11 years. ICD-9 codes for diagnosis/procedures and
NDC codes for medications were extracted from pa-
tients with at least two visits to construct each graph.
Table 1 outlines other statistics about the data.
4.2 Evaluation Tasks and Metrics
The performance of Med2Meta embeddings is
demonstrated using both quantitative and qualitative
evaluations.
4.2.1 Quantitative
The embeddings are evaluated on the following three
medical tasks,
Semantic Similarity Measurement: The seman-
tic similarity between two medical concepts is calcu-
lated as the cosine similarity between their learned
embeddings and Spearman Correlation Coefficient,
ρ, measured against manual rating based on whether
two concepts fall under same hierarchical grouping
of ICD-9 codes collected from clinical classification
software (CCS) (Elixhauser and Palmer, 2015).
Relation Classification: We consider two types
of relation, MAY-TREAT and MAY-PREVENT, from
the National Drug File Reference Terminology (NDF-
RT), which is an ontology containing sets of rela-
tions occurring between drugs and diseases. MAY-
TREAT relation holds for drug-disease pairs where
a drug may be used to treat a disease. While MAY-
PREVENT forms a relation between a drug and a
disease if the drug may be used to prevent the dis-
ease. A 2-nearest neighbor classifier is trained on
relation tuples from triples (rel, c1, c2) where rel ∈
{MAY − T REAT,MAY − PREV ENT }. That is, for
each test tuple, the cosine similarity between the vec-
tor offset between the drug-disease embeddings in-
volved (i.e., m(c1) - m(c2)) and that of all the other tu-
ples in the dataset are first computed. Then the cosine
similarities are ranked in descending order, and the
evaluation metrics (Accuracy, AUC-ROC and AUC-
PR) are measured. If any of the two top-ranked tuples
holds the same relation as the test tuple, it is consid-
ered as a correct match.
Outcome Prediction: This is a binary predic-
tion task that tries to predict whether the patient is
at risk of developing a disease in the future visit,
v
t+1
, trained on visit embedding sequence up to v
t
.
We focus on patients with heart failure (HF) disease.
Thereby, we examine only patients with at least two
visits and check if they contain an occurrence of heart
failure in their v
t+1
visit. These are considered as the
instances belonging to the positive class (HF). As av-
erage number of visits per patient is 2 in MIMIC-III,
v
t+1
in our case is 1. A binary logistic regression clas-
sifier is trained/tested to perform this prediction task.
The train/test/validation split for positive instances is
75%/12.5%/12.5% and the same split is applied to
equal number of total negative instances, where neg-
ative instances are formed with patients who do not
have HF code in their record up to the v
t+1
visit.
4.2.2 Qualitative
We use t-Distributed Stochastic Neighbor Embed-
ding (t-SNE) (Maaten and Hinton, 2008), which is
a visualization technique that maps data in a high-
dimensional space to two or three dimensions, to
examine if natural clusters of medical codes con-
tain similar diagnosis, medication and procedure con-
cepts.
4.3 Baseline Models
To show that the contribution of inclusion of hetero-
geneous data and meta-embedding learning approach
in Med2Meta leads to superior empirical results, we
compare against Med2Meta (M2M) trained on single
view as well as other vector ensemble methods which
include,
M2M d: Proposed approach with embeddings
learned from graph auto-encoder with only demo-
graphics features.
M2M l: Proposed approach with embeddings
learned from graph auto-encoder with only laboratory
test results features.
Med2Meta: Learning Representations of Medical Concepts with Meta-embeddings
373
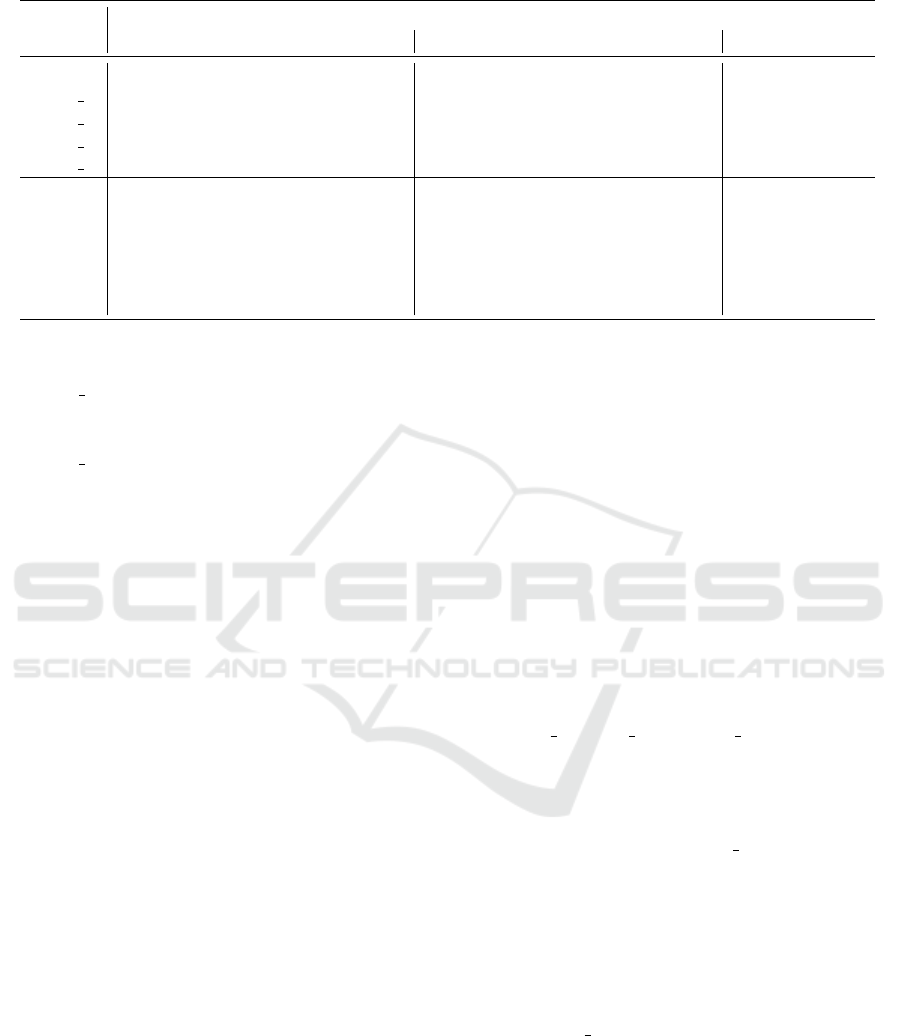

Table 2: Performance of different embeddings on three different tasks.
Heart Failure (HF) Prediction Relation (Rel) Classification Semantic (Sem)
Metrics AUC-ROC Accuracy AUC-PR AUC-ROC Accuracy AUC-PR Similarity ρ
M2M 0.685 0.638 0.674 0.833 0.950 0.967 0.650
M2M d 0.640 0.619 0.635 0.346 0.500 0.692 0.154
M2M l 0.627 0.618 0.629 0.354 0.880 0.923 0.577
M2M n 0.620 0.616 0.630 0.352 0.880 0.750 0.154
M2M s 0.657 0.628 0.632 0.481 0.227 0.385 0.154
CONC 0.666 0.626 0.673 0.481 0.500 0.692 0.154
AVG 0.639 0.627 0.634 0.370 0.700 0.846 0.327
SVD 0.679 0.605 0.673 0.574 0.550 0.615 0.154
Hot 0.520 0.587 0.587 0.426 0.570 0.077 0.119
GV 0.592 0.589 0.575 0.500 0.500 0.692 0.153
M2V 0.678 0.577 0.670 0.648 0.750 0.850 0.576
M2M n: Proposed approach with embeddings
learned from graph auto-encoder with only clinical
notes features.
M2M s: Proposed approach where on the encoder
side instead of dual encoders, only a single encoder is
considered for each view that takes as input the source
embedding, I
i
src
.
CONC: The source feature-specific embeddings
are simply concatenated to represent the final em-
bedding of each medical concept. We `2 normalize
each source feature-specific embedding before con-
catenation to ensure that each embedding contributes
equally during the similarity computation.
AVG: The source feature-specific embeddings are
averaged to represent the final embedding of each
medical concept. `2 normalization is performed on
each source feature-specific embedding before aver-
aging.
SVD: Consider a matrix C of dimension N ×d
svd
,
where d
svd
is dimension of the resulting embedding
from the concatenation of the `
2
normalized embed-
dings for each medical concept. Singular Value De-
composition (SVD) (Golub and Reinsch, 1970) is ap-
plied on C to get the decomposition C = USV
T
. For
each concept, the corresponding vector in U is con-
sidered as the SVD embedding.
We also compare against hot vector representation
and two state-of-the-art embedding models,
Hot: This is the one-hot vector representation of
a concept c, v
c
∈ {0, 1}
N
. Only the dimension corre-
sponding to the concept is set to 1.
GV: GloVe (Pennington et al., 2014) is an un-
supervised learning approach of word embeddings
based on word co-occurrence matrix.
M2V: Med2Vec (Choi et al., 2016a) is a two-layer
neural network for learning lower dimensional repre-
sentations for medical concepts.
5 EXPERIMENTAL RESULTS
5.1 Quantitative Analysis
Table 2 reports results of performance of embeddings
obtained with different models on heart failure (HF)
prediction, semantic similarity (Sem) between med-
ical concepts and relation classification (Rel) tasks.
For heart failure prediction and relation classifica-
tion tasks, the embeddings are evaluated in terms of
AUC-ROC, AUC-PR and Accuracy, and for semantic
similarity with Spearman Correlation Coefficient (ρ).
We see that our multi-view, meta-learning approach
Med2Meta (M2M) outperforms the single view mod-
els M2M d, M2M l and M2M n on all the tasks.
This reinforces the contribution of learning embed-
dings from multi-modal data and means that differ-
ent types of embeddings contribute significantly ac-
cording to their semantic strengths. Among the single
view models, surprisingly, M2M d is seen to perform
better than the other two in HF task. This could be
attributed to most HF patients having distinctive de-
mographics (e.g., older patients).
The benefit of having dual encoders and recon-
structing jointly from the source embedding and av-
erage of most similar medical concepts for each view
can be seen when comparing M2M’s performance
against MSM s, which is ablated version of M2M
with a single encoder for each view. This indicates
that the dual encoder encourages the consolidated fea-
ture space to have locally linear patches for instances
with similar semantics.
Compared to the other types of fusion ap-
proaches, fusing the source embeddings through
meta-embedding learning in M2M leads to consid-
erable AUC-ROC gain on all tasks. AVG perform-
HEALTHINF 2020 - 13th International Conference on Health Informatics
374

(a) Med2Meta (b) CONC (c) AVG (d) SVD
Figure 2: The t-SNE plots of learned embedding spaces. The color of the dot in the Figure indicates the cluster the medical
concept has been assigned to by K-means Clustering.
ing at par with SVD and CONC, aligns with findings
in (Coates and Bollegala, 2018), which found AVG
to outperform them in several benchmark NLP tasks.
With regard to M2M’s superior performance gains,
we can conclude that the joint reconstruction of the
source embedding and its most similar concept em-
bedding aggregation across all the views is able to
integrate both local semantic information in terms of
the closest medical concepts and global semantic in-
formation in terms of multi-modality.
Rows 9-11 in Table 2 show that M2M outperforms
simple representation, one-hot vector, and state-of-
the-art embedding models, GloVe and Med2Vec, on
all the tasks. M2M performs comparably to Med2Vec
in heart failure prediction task and exceptionally by
29% and 13% increases for relation classification
and semantic similarity tasks respectively. Although
Med2Vec also includes demographic information dur-
ing embedding learning, it does not learn from differ-
ent modalities (e.g., lab results, clinical notes) which
can contain salient information for predictive tasks -
as is confirmed by better performance of M2M across
all the tasks.
5.2 Qualitative Analysis
Our embeddings are qualitatively assessed by visual-
izing their t-SNE plots in 2-D space shown in Figure
2a. All the medical concepts are first grouped into
clusters using K-means Clustering. The color of the
dot in Figures 2a–2d indicates the cluster the medical
concept has been assigned to. For comparison, t-SNE
plots of embeddings obtained by the baselines CONC,
SVD and AVG are also shown in Figures 2b, 2c, 2d. It
is evident that Med2Meta has been able to learn med-
ically meaningful representations such that they have
been separated into distinct clusters compared to the
baseline plots.
6 RELATED WORKS
Meta-embedding Learning: Usefulness of meta-
embeddings in NLP tasks are realized in some very
recent works. (Yin and Sch
¨
utze, 2015) is one of
the first works in meta-embedding learning that pro-
posed a model, 1TON, that learns meta-embeddings
by projecting them to source embeddings using sep-
arate projection matrices. 1TON is then extended
to 1TON+ to account for out-of-vocabulary (OOV)
words by first predicting their source embeddings. An
unsupervised locally-linear approach is used in (Bol-
legala et al., 2017) to learn meta-embedding of each
word based on its local neighborhood of word embed-
dings. Different variants of autoencoder are used in
(Bollegala and Bao, 2018) to learn meta-embeddings
from pre-trained word embeddings.
Some other works did not learn meta-embeddings,
but closely resemble its mechanism by demonstrat-
ing effectiveness of integration of different types of
embeddings. (Ma et al., 2018) learns similarities
between drugs by integrating embeddings learned
from a multi-view graph auto-encoder using attention
mechanism. A two-sided neural network is used in
(Luo et al., 2014) to learn embeddings from multiple
data sources.
7 CONCLUSION
Leveraging the heterogeneous data types in EHR can
be beneficial to learning embeddings that holistically
reflect all semantic properties among different med-
ical concepts. Our proposed approach, Med2Meta,
learns feature-specific embeddings using a graph
auto-encoder by considering each data type as a sepa-
rate view. It then models integration of embeddings
as a meta-embedding learning problem so that la-
tent similarities and natural clusters between medical
concepts are captured in the meta-embedding space
through joint reconstruction across all the views. Em-
pirical results on three different tasks and visualiza-
tion with t-sne plots establish the superior perfor-
mance and efficacy of Med2Meta over baselines.
REFERENCES
Bengio, Y., Courville, A., and Vincent, P. (2013). Represen-
tation learning: A review and new perspectives. IEEE
Med2Meta: Learning Representations of Medical Concepts with Meta-embeddings
375

transactions on pattern analysis and machine intelli-
gence, 35(8):1798–1828.
Bollegala, D. and Bao, C. (2018). Learning word meta-
embeddings by autoencoding. In Proceedings of the
27th International Conference on Computational Lin-
guistics, pages 1650–1661.
Bollegala, D., Hayashi, K., and Kawarabayashi, K.-i.
(2017). Think globally, embed locally—locally
linear meta-embedding of words. arXiv preprint
arXiv:1709.06671.
Charles, D., Gabriel, M., and Furukawa, M. F. (2013).
Adoption of electronic health record systems among
us non-federal acute care hospitals: 2008-2012. ONC
data brief, 9:1–9.
Chen, Y., Perozzi, B., Al-Rfou, R., and Skiena, S. (2013).
The expressive power of word embeddings. arXiv
preprint arXiv:1301.3226.
Choi, E., Bahadori, M. T., Searles, E., Coffey, C., Thomp-
son, M., Bost, J., Tejedor-Sojo, J., and Sun, J. (2016a).
Multi-layer representation learning for medical con-
cepts. In Proceedings of the 22nd ACM SIGKDD In-
ternational Conference on Knowledge Discovery and
Data Mining, pages 1495–1504. ACM.
Choi, E., Bahadori, M. T., Song, L., Stewart, W. F., and
Sun, J. (2017). Gram: graph-based attention model for
healthcare representation learning. In Proceedings of
the 23rd ACM SIGKDD International Conference on
Knowledge Discovery and Data Mining, pages 787–
795. ACM.
Choi, E., Schuetz, A., Stewart, W. F., and Sun, J. (2016b).
Medical concept representation learning from elec-
tronic health records and its application on heart fail-
ure prediction. arXiv preprint arXiv:1602.03686.
Choi, Y., Chiu, C. Y.-I., and Sontag, D. (2016c). Learning
low-dimensional representations of medical concepts.
AMIA Summits on Translational Science Proceedings,
2016:41.
Chowdhury, S., Zhang, C., Yu, P. S., and Luo, Y. (2019).
Mixed pooling multi-view attention autoencoder for
representation learning in healthcare. arXiv preprint
arXiv:1910.06456.
Coates, J. and Bollegala, D. (2018). Frustratingly easy
meta-embedding–computing meta-embeddings by av-
eraging source word embeddings. arXiv preprint
arXiv:1804.05262.
Elixhauser, A. and Palmer, L. (2015). Clinical classifica-
tions software (ccs): Agency for healthcare research
and quality; 2014 [cited 2015].
Goldberg, Y. (2016). A primer on neural network models
for natural language processing. Journal of Artificial
Intelligence Research, 57:345–420.
Golub, G. H. and Reinsch, C. (1970). Singular value de-
composition and least squares solutions. Numerische
mathematik, 14(5):403–420.
Jensen, P. B., Jensen, L. J., and Brunak, S. (2012). Mining
electronic health records: towards better research ap-
plications and clinical care. Nature Reviews Genetics,
13(6):395.
Johnson, A. E., Pollard, T. J., Shen, L., Li-wei, H. L.,
Feng, M., Ghassemi, M., Moody, B., Szolovits, P.,
Celi, L. A., and Mark, R. G. (2016). Mimic-iii, a
freely accessible critical care database. Scientific data,
3:160035.
Knake, L. A., Ahuja, M., McDonald, E. L., Ryckman,
K. K., Weathers, N., Burstain, T., Dagle, J. M., Mur-
ray, J. C., and Nadkarni, P. (2016). Quality of ehr data
extractions for studies of preterm birth in a tertiary
care center: guidelines for obtaining reliable data.
BMC pediatrics, 16(1):59.
Luo, Y., Tang, J., Yan, J., Xu, C., and Chen, Z. (2014).
Pre-trained multi-view word embedding using two-
side neural network. In AAAI, pages 1982–1988.
Ma, T., Xiao, C., Zhou, J., and Wang, F. (2018). Drug sim-
ilarity integration through attentive multi-view graph
auto-encoders. arXiv preprint arXiv:1804.10850.
Maaten, L. v. d. and Hinton, G. (2008). Visualizing data
using t-sne. Journal of machine learning research,
9(Nov):2579–2605.
Miotto, R., Li, L., Kidd, B. A., and Dudley, J. T. (2016).
Deep patient: an unsupervised representation to pre-
dict the future of patients from the electronic health
records. Scientific reports, 6:26094.
Nair, V. and Hinton, G. E. (2010). Rectified linear units
improve restricted boltzmann machines. In Proceed-
ings of the 27th international conference on machine
learning (ICML-10), pages 807–814.
Pennington, J., Socher, R., and Manning, C. (2014). Glove:
Global vectors for word representation. In Proceed-
ings of the 2014 conference on empirical methods in
natural language processing (EMNLP), pages 1532–
1543.
Roweis, S. T. and Saul, L. K. (2000). Nonlinear dimension-
ality reduction by locally linear embedding. science,
290(5500):2323–2326.
Shickel, B., Tighe, P. J., Bihorac, A., and Rashidi, P. (2018).
Deep ehr: A survey of recent advances in deep learn-
ing techniques for electronic health record (ehr) anal-
ysis. IEEE journal of biomedical and health informat-
ics, 22(5):1589–1604.
Tran, T., Nguyen, T. D., Phung, D., and Venkatesh, S.
(2015). Learning vector representation of medical ob-
jects via emr-driven nonnegative restricted boltzmann
machines (enrbm). Journal of biomedical informatics,
54:96–105.
Wold, S., Esbensen, K., and Geladi, P. (1987). Principal
component analysis. Chemometrics and intelligent
laboratory systems, 2(1-3):37–52.
Yin, W. and Sch
¨
utze, H. (2015). Learning meta-embeddings
by using ensembles of embedding sets. arXiv preprint
arXiv:1508.04257.
HEALTHINF 2020 - 13th International Conference on Health Informatics
376
