
A MEDICAL INFORMATION SYSTEM TO MANAGE A CANCER
DATABASE
André Cid Ferrizzi, Toni Jardini, Leandro Rincon Costa, Jucimara Colombo
Paula Rahal, Carlos Roberto Valêncio
UNESP, São José do Rio Preto, São Paulo, Brazil
Edmundo Carvalho Mauad, Lígia Maria Kerr, Geraldo Santiago Hidalgo
Hospital do Câncer de Barretos, Barretos, São Paulo, Brazil
Keywords: Medical Information System, Cancer Database, Bioinformatics.
Abstract: Cancer is the second main cause of death in Brazil, and according to statistics disclosed by INCA – National
Cancer Institute 466,730 new cases of the disease are forecast for 2008. The storage and analysis of tumour
tissues of various types and patients' clinical data, genetic profiles, characteristics of diseases and
epidemiological data may provide more precise diagnoses, providing more effective treatments with higher
chances for the cure of cancer. In this paper we present a Web system with a client-server architecture,
which manages a relational database containing all information relating to the tumour tissue and their
location in freezers, patients, medical forms, physicians, users, and others. Furthermore, it is also discussed
the software engineering used to developing the system.
1 INTRODUCTION
Cancer is a fairly diverse disease and the multiple
genetic and epigenetic changes peculiar to it make
its prevention, diagnosis and therapy difficult. Thus,
studies that are meant to establish the tumour’s
molecular genetic profile are essential so that one
can understand the disease's complexity, establish
the biological basis and provide means to identify
the best therapeutic strategies, since, in spite of the
developments in chemotherapy, in surgical
techniques and in drug combinations, there are types
of neoplasies in which there has been practically no
prognosis improvement within the last ten years, a
fact that underscores the need to know the tumour's
biology (O’Connor, 2007) (He et al., 2007).
Cancer is the second main cause of death in
Brazil, and according to statistics disclosed by INCA
– National Cancer Institute 466,730 new cases of the
disease are forecast for 2008. Out of these, 231,860
are expected to be new male patient cases, while
234,870 are expected to be new female patient cases
(Brazilian National Cancer Institute, 2008).
The Barretos Cancer Hospital - SP is showcase
for a cancer treatment hospital. It rates among the
largest cancer hospitals in Brazil. Over 400 thousand
consultations take place and over 6 thousand new
cases of cancer, from all Brazilian states, are
diagnosed at the facility, every year. Since the
Hospital treats patients from a number of states and
regions within Brazil, the Barretos Cancer Hospital’s
Information System to Manage the Tumour Bank
(SCGBT) will make it possible to develop studies in
the area of prognosis, diagnosis and therapeutic
markers in representative samples of the Brazilian
population.
This article is meant to describe the SCGBT, the
system that underpins the whole management
process of a tumour samples database, by showing
its architecture, construction processes,
functionalities, and presenting some extracted data.
2 METHODOLOGY
The software development process employed is the
Unified Process – UP (Larman, 2001); Modeling of
some aspects of the system was performed with the
Unified Modeling Language (UML) and for
configuration management, Subversion (SVN) is
used (Subversion, 2008).
268
Cid Ferrizzi A., Jardini T., Rincon Costa L., Colombo J., Rahal P., Roberto Valêncio C., Carvalho Mauad E., Maria Kerr L. and Santiago Hidalgo G.
(2008).
A MEDICAL INFORMATION SYSTEM TO MANAGE A CANCER DATABASE.
In Proceedings of the Third International Conference on Software and Data Technologies - ISDM/ABF, pages 268-271
DOI: 10.5220/0001881802680271
Copyright
c
SciTePress
The programming language employed is PHP
Hypertext Preprocessor (PHP) and he Database
Management System (DBMS) used was MySQL.
3 SCGBT ENGINEERING
The whole SCGBT implementation was supported
by Project Management Body of Knowledge
(PMBOK) project management models and
processes, focused on four main areas: Requirements
Management, Configurations Management, Risks
Management and Tests Management (Project
Management Institute, 2004).
4 DATABASE MANAGED BY
SCGBT
The Barretos Cancer Hospital database comprises
information provided by the system’s users, patients,
doctors, tissue samples, medical forms, freezers,
departments, among others. It is possible to use data
mining, like some techniques discussed by Han &
Kamber (2006), to obtain knowledge in order to
diagnose cancer-related diseases.
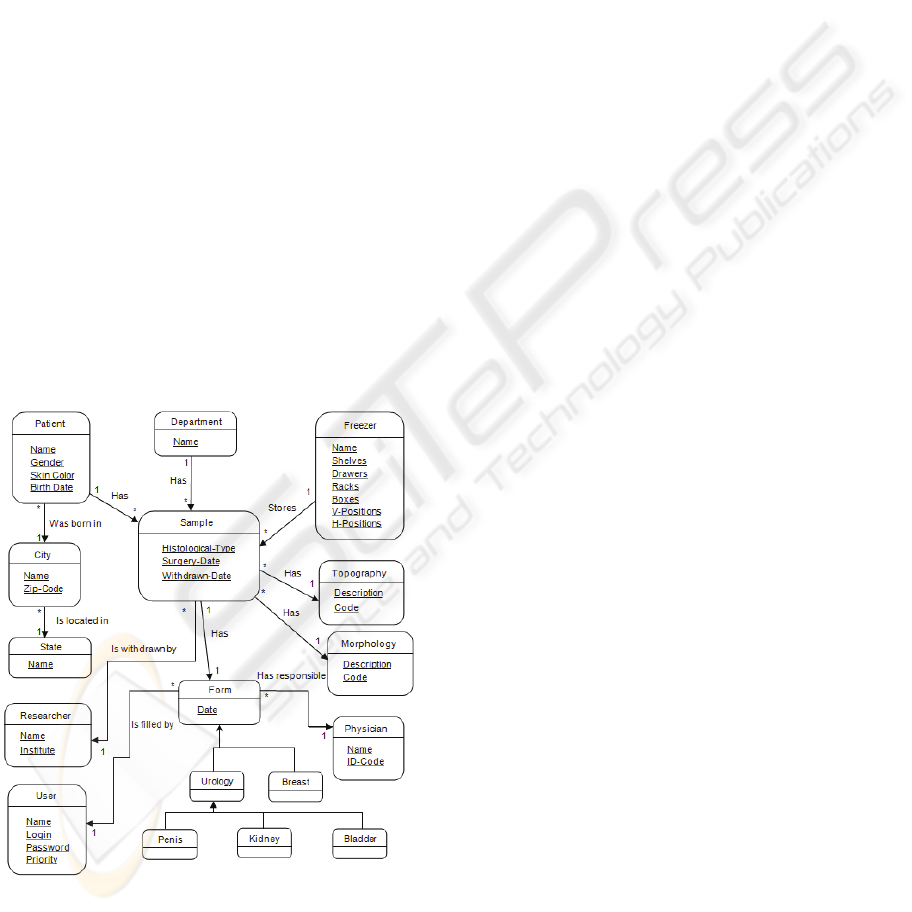
Figure 1: Part of the HCB database class diagram.
Fig. 1 shows a part of the database UML class
diagram. In this diagram, the main items of the
database are shown:
Patient: represents the patient treated at the
Hospital;
Sample: it is a tumour sample collected from
the patient; this may be of the following types:
normal, tumour, blood, leukocytes, DNA,
RNA and serum;
Topography: the International Classification of
Diseases for Oncology Code (ICD-O) is used
(Pan-American Health Organization, 2005);
Morphology: the ICD-O is used for this
attribute's values;
Freezer: is the location where the samples are
stored;
Researcher: a sample may be removed from the
freezer for research purposes;
Forms: show information on the patient's
history, diagnosis, clinical state, treatment and
prognosis;
Doctor: is the doctor in charge of the patient.
5 SCGBT FUNCTIONALITIES
SGCBT comprises a number of functionalities,
providing full handling of the data on each patient
on record, as well as the tumour samples and their
characteristics. The next sections show the main
SCGBT's functionalities.
5.1 Patients
SCGBT comprises functionalities that handle patient
data, such as inclusion, update, removal and
viewing. In order to retrieve these data, the system
offers an interactive interface for the definition of
searches by the users, known as patient filtering. By
means of this interface, the user is able to put
together his/her patient search according to certain
parameters. The user is able to add or remove search
parameters, which include sample location,
topography, morphology, patient’s age, type of
sample, name, among others.
5.2 Samples
To handle the tumour tissue samples data, add,
remove, change and view samples functionalities are
available. The samples’ morphology and topography
fields use ICD-O, which associates a code and a
description to each morphology and topography.
A MEDICAL INFORMATION SYSTEM TO MANAGE A CANCER DATABASE
269
5.3 Forms
Each sample is associated to a single patient. For
each one of these samples, it is possible to fill out,
update and view its relevant forms. Such forms hold
data on the tumour's identification and history,
diagnosis, clinical stage, treatment and prognosis.
All forms stored in the database are easily viewed
and changed as required.
5.4 Freezers, Users and Doctors
It is possible to enter new freezers on record,
specifying its physical dimensions such as number
of shelves, racks, drawers, boxes and positions. The
system also offers the possibility of handling these
data. Thus, operations such as freezer searches and
changes can be easily performed. In order to
guarantee security and access control, SCGBT
encompasses user management, allowing the control
of users entered in the system, by means of
privileges. There is also information management
with regard to the doctors in charge of the each
patient’s diagnosis, who are associated with various
departments within the Hospital.
5.5 Reports
SCGBT makes several types of reports available in
order to extract data from the cancer database.
Several types of sample and patient reports are
contemplated. Data collection and management, by
means of our computer system, began in 2006 and at
the present juncture, after about two years’ time, the
Tumour Bank has almost ten thousand samples from
almost three thousand patients, 1383 (52.55%) of
whom are male, and 1249 (47.45%) are female.
With regard to the samples, 2306 are normal tissue
samples, 3221 are tumour tissue samples, 353 are
serum-type samples, 325 are leukocyte-type samples
and 3698 are blood-type samples. Fig. 2 shows some
data collected from one of the patient sample
reports.
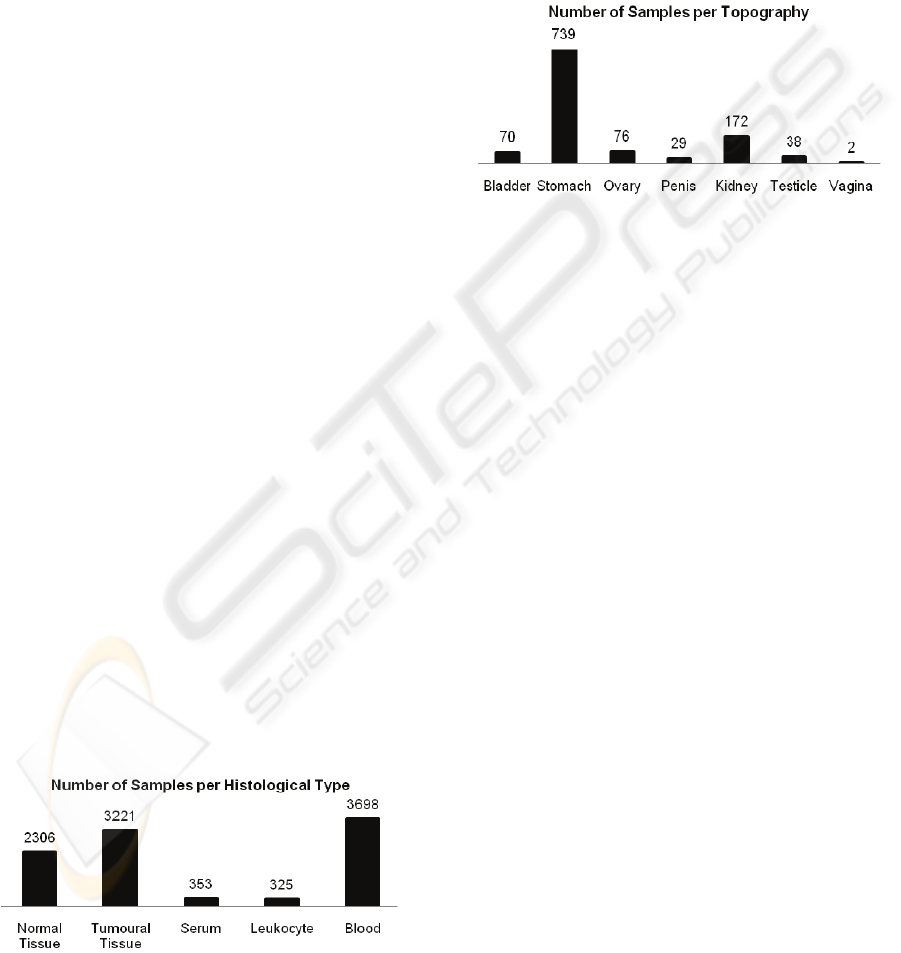
Figure 2: Number of samples per histological type.
The forms are yet to be filled out in their entirety
since they are in the implementation stage. At the
present juncture, there are thirteen forms filled out
on penis cancer, fifty-eight on kidney cancer and
twenty-one on bladder cancer. Forms are established
in order to provide data to publish papers on cancer,
such as the paper on penis cancer (Babeto et al.,
2007). Fig. 3 provides some data on the number of
tumour samples collected per patient’s organ.
Figure 3: Number of tumour tissue samples per
topography.
6 THE BARRETOS CANCER
HOSPITAL AND THE AC
CAMARGO HOSPITAL IN SÃO
PAULO DATABASES
INTEGRATION
An important segment of this project is the
integration of the Barretos Cancer Hospital SCGBT
with the central bio-repository kept by the A.C.
Camargo Cancer Hospital in São Paulo (AC
Camargo Cancer Hospital, 2008). This bio-
repository is meant to receive and maintain, in a
centralized manner, the data from the associated
cancer research centers. By means of a data
communication system, Barretos Cancer Hospital
will provide its relevant data to this bio-repository.
At the present juncture, the data transmission model
is being prepared. Its basis will be the YAML data
structuring language (YAML Ain’t Markup
Language), a language that is similar to the widely-
known XML, but which is easier for humans to read
and understand (YAML, 2008).
7 CONCLUSIONS
The purpose of this paper is to introduce SCGBT,
which manages a cancer database. The use of human
tissue in the study is vital, since within the last few
ICSOFT 2008 - International Conference on Software and Data Technologies
270

decades there has been a decrease in the use of
animal cellular lineage and models in the study of
cancer. This trend has taken place concurrently with
the development of molecular studies and also with
the conception of a the neoplasic phenomenon as a
heterotypical process in which both the neoplasic
cell and the issue environment in which it develops
play a key role, since, in addition to the genetic
factors associated with the tumour, the individual-
related factors interfere with the tumour and its
response to treatment (Marahatta, 2005).
8 FUTURE WORK
As future work the integration with other tumour
banks will be developed. Another task to be
performed is the inclusion of reports and charts that
may provide a view of the distribution of the various
cancer-related data by geographic region (Carr et al.,
2003).
ACKNOWLEDGEMENTS
We wish to express our appreciation to FAPESP and
PROPG (Pró-reitoria de pós-graduação) for
providing financial support for this work; to
Fundação Pio XII, to IBILCE – UNESP, to Tamara
Colaiacovo for their collaboration to this project and
Dr. Geraldo Santiago Hidalgo, pathologist
responsible for the beginning of this project.
REFERENCES
AC Camargo Cancer Hospital, 2008. Available at: http://
http://www.hcanc.org.br (accessed 21 April 2008).
Babeto, E., Pires, L. C., Valsechi, M. C., Ferrizzi, A. C.,
Valencio, C. R., Kerr, L. M., Faria, E. F., Seabra, D.,
Soares, F. A., Peitl, P. J., Rahal, P., 2007. Differential
Gene Expression Analysis in Penile Carcinoma, In
VIII São Paulo Research Conference: "Câncer 2007:
da Biologia Molecular ao Tratamento".
Brazilian National Cancer Institute, 2008. Available at:
http://www.inca.gov.br (accessed 21 April 2008).
Carr, D. B., Bell, S., Pickle, L., Zhang, Y., Li, Y., 2003.
The State Cancer Profiles Web Site and Extensions of
Linked Micromap Plots and Conditioned Choropleth
Map Plots. In Proceedings of the 2003 Annual
National Conference on Digital Government
Research, Boston.
Han, J. & Kamber, M., 2006. Data Mining: Concepts and
Techniques. Morgan Kaufmann Publisher, São
Francisco.
He, M., et al., 2007. Cancer Development and
Progression. Adv Exp Med Biol, vol. 593, pp.117-
133.
Larman, C., 2001. Applying UML and Patterns: An
Introduction to Object-Oriented Analysis and Design
and Iterative Development. Bookman, 2
nd
Ed.
Marahatta, S. B., 2005. Cancer: Determinants and
Progression. Nepal Med Coll J. vol.7, pp. 65-71.
O’Connor, R., 2007. The Pharmacology of Cancer
Resistance. Anticancer Res, vol. 27, pp.1267-1272.
Pan-American Health Organization, 2005. World Health
Organization. ICD-O: International Classification of
Diseases for Oncology, EDUSP, Portuguese Edition.
Project Management Institute, 2004. A Guide to the
Project Management Body of Knowledge: PMBOK
Guide, Project Management Institute, 3
rd
Ed.
Subversion, 2008. Available at: http://subversion.tigris.org
(accessed 21 April 2008).
YAML, 2008. Available at: http://www.yaml.org (accessed
21 April 2008).
A MEDICAL INFORMATION SYSTEM TO MANAGE A CANCER DATABASE
271
