
SynapCountJ: A Tool for Analyzing Synaptic Densities in Neurons
Gadea Mata
1
, J
´
onathan Heras
1
, Miguel Morales
2
, Ana Romero
1
and Julio Rubio
1
1
Departamento de Matem
´
aticas y Computaci
´
on, Universidad de La Rioja, Logro
˜
no, La Rioja, Spain
2
Institut de Neurocincies, Universitat Aut
`
onoma de Barcelona, Barcelona, Spain
Keywords:
Synapses, Synaptic Density, Image Processing, ImageJ.
Abstract:
The quantification of synapses is instrumental to measure the evolution of synaptic densities of neurons under
the effect of some physiological conditions, neuronal diseases or even drug treatments. However, the manual
quantification of synapses is a tedious, error-prone, time-consuming and subjective task; therefore, tools that
might automate this process are desirable. In this paper, we present SynapCountJ, an ImageJ plugin, that
can measure synaptic density of individual neurons obtained by immunofluorescence techniques, and also
can be applied for batch processing of neurons that have been obtained in the same experiment or using the
same setting. The procedure to quantify synapses implemented in SynapCountJ is based on the colocalization
of three images of the same neuron (the neuron marked with two antibody markers and the structure of the
neuron) and is inspired by methods coming from Computational Algebraic Topology. SynapCountJ provides
a procedure to semi-automatically quantify the number of synapses of neuron cultures; as a result, the time
required for such an analysis is greatly reduced.
1 INTRODUCTION
Synapses are the points of connection between neu-
rons, and they are dynamic structures subject to
a continuous process of formation and elimination.
Pathological conditions, such as the Alzheimer dis-
ease, have been related to synapse loss associated
with memory impairments. Hence, the possibility
of changing the number of synapses may be an im-
portant asset to treat neurological diseases (Selkoe,
2002). To this aim, it is necessary to determine the
evolution of synaptic densities of neurons under the
effect of some physiological conditions, neuronal dis-
eases or even drug treatments.
The procedure to quantify synaptic density of a
neuron is usually based on the colocalization be-
tween the signals generated by two antibodies (Cuesto
et al., 2011). Namely, neuron cultures are permeabi-
lized and treated with two different primary markers
(for instance, bassoon and synapsin). These antibod-
ies recognize specifically two presynaptic structures.
Then, it is necessary a secondary antibody couple at-
tached to different fluorochromes (for instance red
and green; note, that several other combinations of
color are possible) making these two synaptic proteins
visible under the fluorescence microscope. The two
markers are photographed in two gray-scale images;
that, in turn, are overlapped using respectively the red
and green channels. In the resultant image, the yel-
low points (colocalization of the code channels) are
the candidates to be the synapses.
The final step in the above procedure is the selec-
tion of the yellow points that are localized either on
the dendrites of the neuron or adjacent to them. Tools
like MetaMorph (Devices, 2015) or ImageJ (Schnei-
der et al., 2012) — a Java platform for image pro-
cessing that can be easily extended by means of plu-
gins — can be used to manually count the number of
synapses; however, such a manual quantification is a
tedious, time-consuming, error-prone, and subjective
task; hence, tools that might automate this process
are desirable. In this paper, we present SynapCountJ,
an ImageJ plugin, that semi-automatically quantifies
synapses and synaptic densities in neuron cultures.
2 METHODOLOGY
SynapCountJ supports two execution modes: individ-
ual treatment of a neuron and batch processing — the
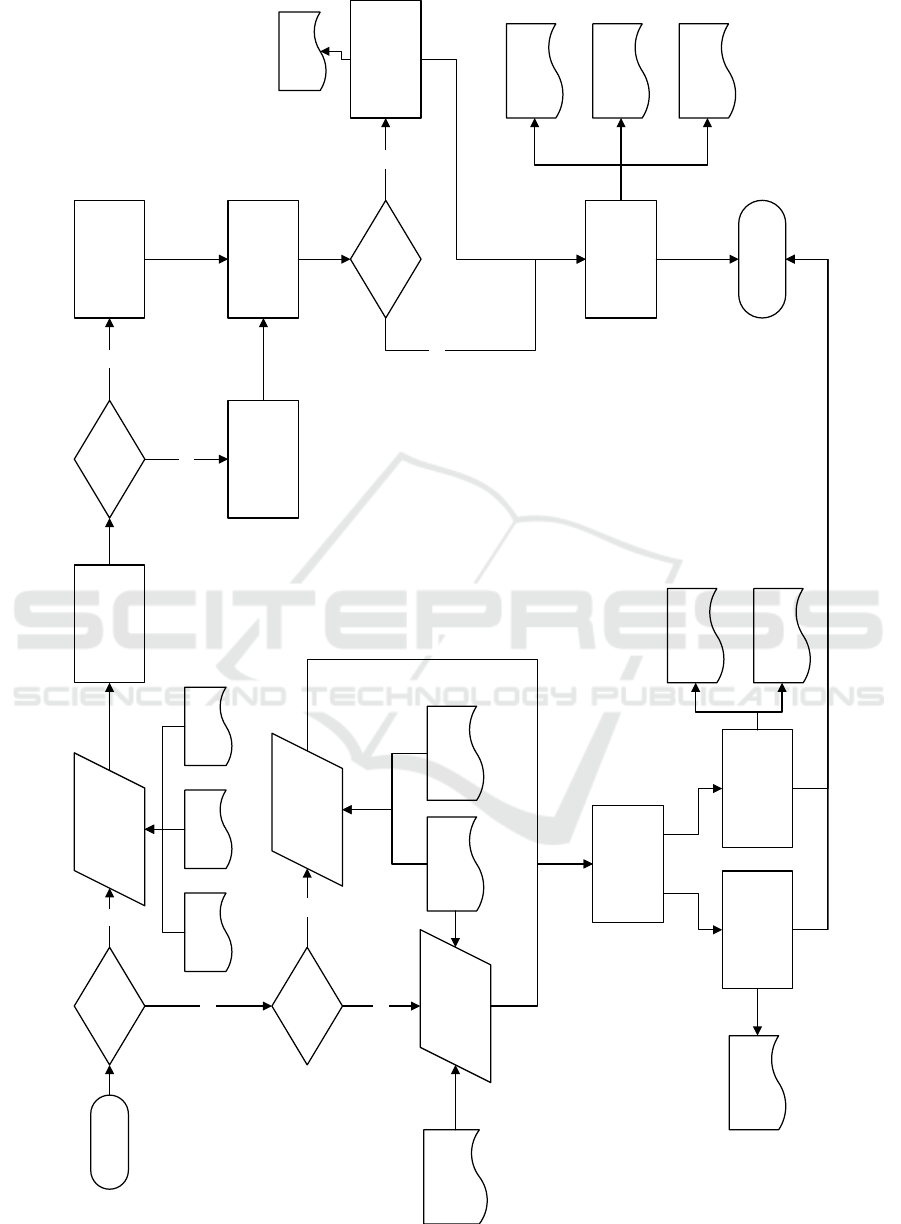
workflow of both modes is provided in Figure 1
Mata, G., Heras, J., Morales, M., Romero, A. and Rubio, J.
SynapCountJ: A Tool for Analyzing Synaptic Densities in Neurons.
DOI: 10.5220/0005637700250031
In Proceedings of the 9th International Joint Conference on Biomedical Engineering Systems and Technologies (BIOSTEC 2016) - Volume 2: BIOIMAGING, pages 25-31
ISBN: 978-989-758-170-0
Copyright
c
2016 by SCITEPRESS – Science and Technology Publications, Lda. All rights reserved
25
Start
Batch
processing?
NO Configure setting
Channel
Green Image
Channel Red
Image
Structure
(txt-file)
Known
threshold?
Select thresholdNO
Write threshold
YES
Save the
settings?
Image with
analyzed region
Image with
indicated
synapses
Xml-file with
the settings
YES
Show the results
Table with
analysis
YES
Path of
directory with
images
NO
Lif-files?
Xml-file with
settings
YES
Lif-file
End
Calculate number
of synapses and
density
Show table Save results images
Table with
analysis
Image with
analyzed region
Image with
indicated
synapses
NO
Save settings
Calculate number of
synapses and
density
Choose input
files
Input path of
files
Choose
directories
Figure 1: Workflow of SynapCountJ.
BIOIMAGING 2016 - 3rd International Conference on Bioimaging
26
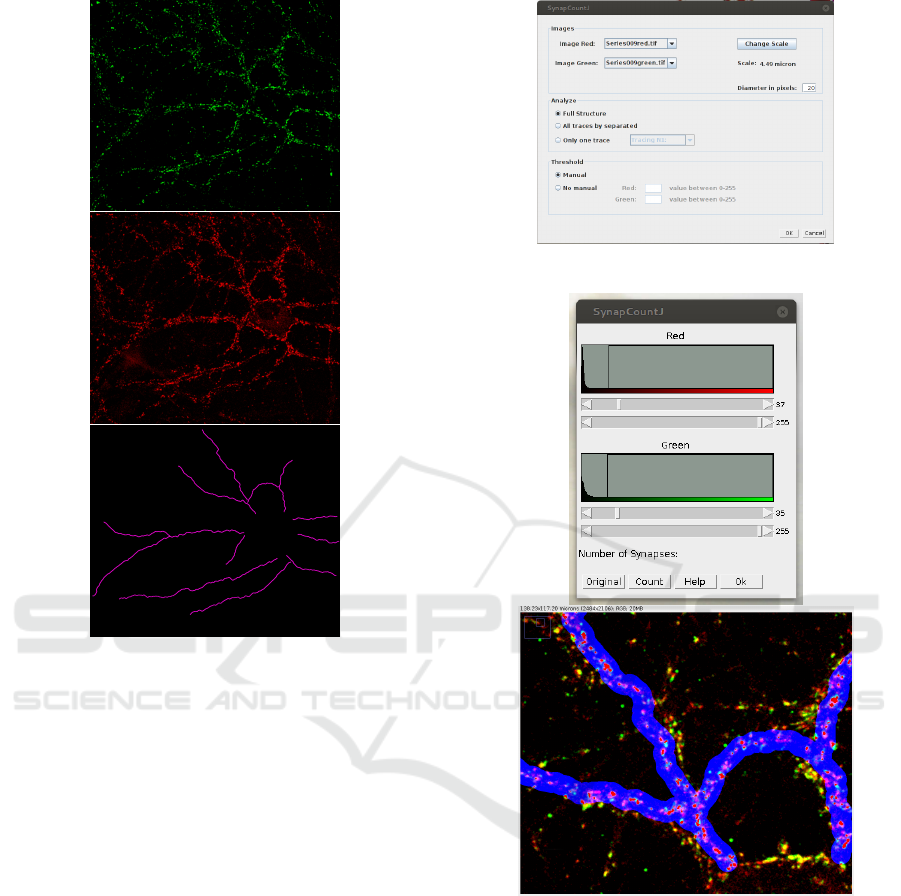
Figure 2: Neuron with two antibody markers and its struc-
ture. Top. Neuron marked with the bassoon antibody
marker. Center. Neuron marked with the synapsin antibody
marker. Bottom. Structure of the neuron.
2.1 Individual Treatment of a Neuron
The input of SynapCountJ in this execution mode are
two images of a neuron marked with two antibodies
(an image per antibody), see Figure 2. SynapCountJ
is able to read tiff (a standard format for biological im-
ages) and lif files (obtained from Leica confocal mi-
croscopes) — the latter requires the Bio-Formats plu-
gin (Linkert et al., 2010). The following steps are ap-
plied to quantify the number of synapses in the given
images.
In the first step, from one of the two images, the
region of interest (i.e. the dendrites where the quan-
tification of synapses will be performed) is specified
using NeuronJ (Meijering et al., 2004) — an ImageJ
plugin for tracing elongated image structures. In this
way, the background of the image is removed. The
result is a file containing the traces of each dendrite
of the image.
Subsequently, the user can decide whether she
wants to perform a global analysis of the whole neu-
ron, or a local analysis focused on each dendrite of
Figure 3: SynapCountJ window to configure the analysis.
Figure 4: SynapCountJ window to modify the threshold of
the red and green channels. Top. Window to fix the thresh-
old of the image. Bottom. Fragment of the neuron image
with the synapses indicated as the red areas on the structure
of the neuron marked in blue. Moving the scrollbars of left
window, the marked areas of the image are changed.
the neuron. In both cases, SynapCountJ requires ad-
ditional information such as the scale and the mean
thickness (that is determined by the size of the subja-
cent dendrite) of the region to analyze (see Figure 3)
— these parameters determine the area of the dendrite
avoiding the background (i.e. all the non-synaptic
marking).
Taking into account the settings provided by the
user, SynapCountJ overlaps the two original images
SynapCountJ: A Tool for Analyzing Synaptic Densities in Neurons
27
of the neuron and the structure of the neuron pre-
viously defined. From the resultant image, Synap-
CountJ identifies the almost white points (the result
of green, red, and blue combination) as synaptic can-
didates, and it allows the user to modify the values
of the red and green channels in order to modify the
detection threshold (see Figure 4).
Once that the detection threshold has been fixed,
the counting process is started. Such a process is
inspired by techniques coming Computational Alge-
braic Topology. In spite of being an abstract math-
ematical subject, Algebraic Topology has been suc-
cessfully applied in digital image analysis (S
´
egonne
et al., 2003; Gonz
´
alez-D
´
ıaz and Real, 2005) In our
particular case, the white areas are segmented from
the overlapped image, and the colors of the resultant
image are inverted — obtaining as a result a black-
and-white image where the synapses are the black ar-
eas. From such an image, the problem of quantifying
the number of synapses is reduced to compute the ho-
mology group in dimension 0 of the image; this corre-
sponds to the computation of the number of connected
components of the image.
Finally, SynapCountJ returns a table with the ob-
tained data (length of dendrites both in pixels and mi-
cras, number of synapses, and density of synapses per
100 micron) and two images showing, respectively,
the analyzed region and the marked synapses (see Fig-
ure 5).
2.2 Batch Processing
Images obtained from the same biological experiment
usually have similar settings; hence, their processing
in SynapCountJ will use the same configuration pa-
rameters. In order to deal with this situation, Synap-
CountJ can be applied for batch processing of several
images using a configuration file. It is necessary to
study at least one image from experiment to get the
optimal settings. The parameters are saved and used
to process the set of images from the same experi-
ment.
For batch processing, SynapCountJ reads tiff files
organized in folders or a lif file (the kind of files pro-
duced by Leica confocal microscopes), and using the
configuration file processes the different images. As
a result, a table with the information related to each
neuron from the batch (the table includes an analy-
sis for both the whole neuron and from each of its
dendrites) is obtained. In addition, in the same direc-
tory where the lif-file or tiff-files are stored, the plu-
gin saves all the resultant images for each image from
experiment (one of them shows the marked synapses
and the other one, the region which has been studied).
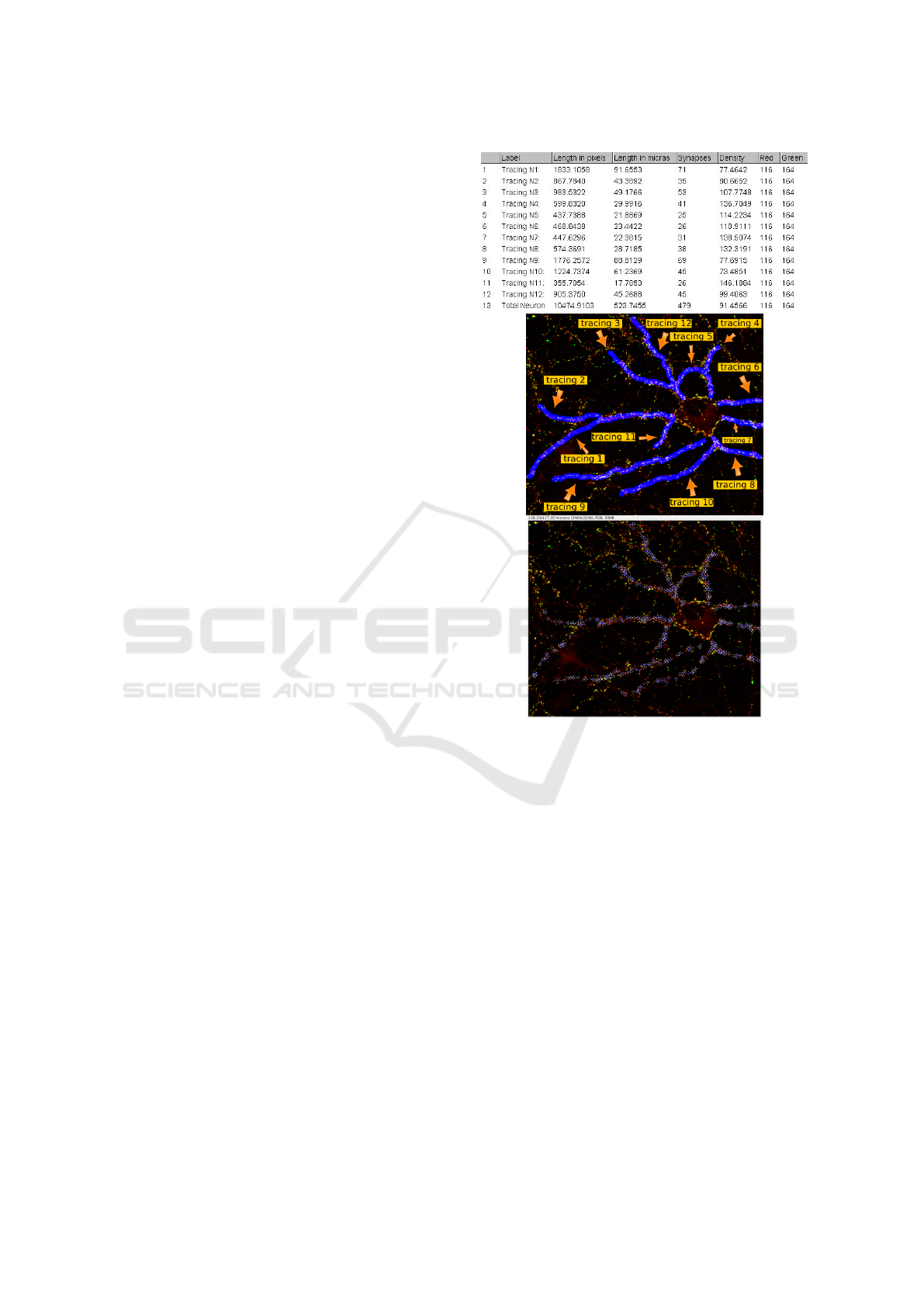
Figure 5: Results provided by SynapCountJ. Top. Table
with the results obtained by SynapCountJ. Center. Image
with the analyzed region of the neuron. Bottom. Image with
the counted synapses indicated by means of blue crosses.
3 EXPERIMENTAL RESULTS
The original aim of SynapCountJ was the automatic
analysis of synaptic density on neurons treated with
SB 415286 — an organic inhibitor of GSK3, a ki-
nase which inhibition was proposed as a therapy in
AD treatment (DaRocha-Souto et al., 2003) — such
a treatment, as it was previously demonstrated, pro-
motes synaptogenesis and spinogenesis in primary
cultures of rodent hippocampal neurons and in Dros-
ophyla neurons (Cuesto et al., 2015; Franco et al.,
2004). In this setting, a comparative study has been
performed in order to evaluate the results that can be
obtained with SynapCountJ.
Primary hippocampal cultures were obtained from
P0 rat pups (Sprague-Dawley, strain, Harlan Labo-
BIOIMAGING 2016 - 3rd International Conference on Bioimaging
28
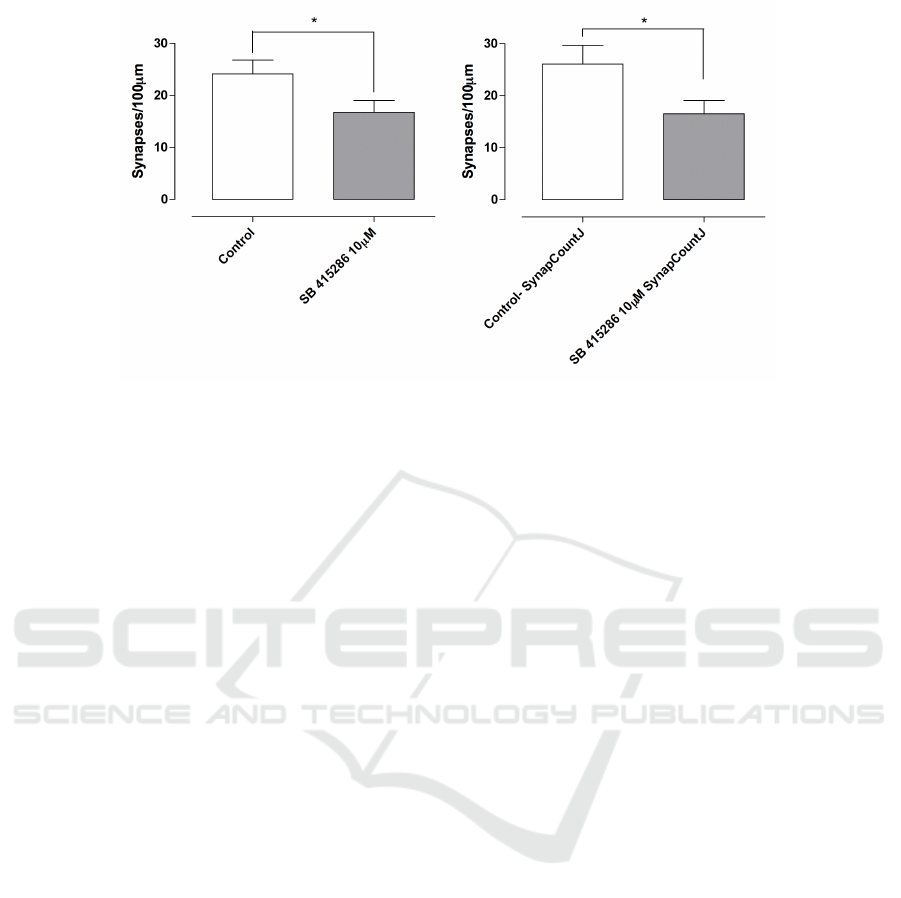
Figure 6: Quantification of synapses. Left. Manual quantification of synapses. Right. Quantification of synapses using
SynapCountJ.
ratories Models SL, France). Animals were anes-
thetized by hypothermia in paper-lined towel over
crushed-ice surface during 2-4 minutes and eutha-
nized by decapitation. Animals were handled and
maintained in accordance with the Council Direc-
tive guidelines 2010/63EU of the European Parlia-
ment. Briefly, glass coverslips (12 mm in diame-
ter) were coated with poly-L-lysine and laminin, 100
and 4 µg/ml respectively. Neurons at a 10 × 104
neurons/cm
2
density were seeded and grown in Neu-
robasal (Invitrogen, USA) culture medium supple-
mented with glutamine 0.5 mM, 50 mg/ml penicillin,
50 units/ml streptomycin, 4% FBS and 4% B27 (In-
vitrogen, CA, USA), as described before in (Cuesto
et al., 2011). At days 4, 7 and 14 in culture a 20%
of culture medium was replace by fresh medium.
Cytosine-D-arabinofuranoside (4 µM) was added to
prevent overgrowth of glial cells (day 4).
Synaptic density on hippocampal cultures was
identified as previously described in (Cuesto et al.,
2011). In short, cultures were rinsed in phosphate
buffer saline (PBS) and fixed for 30 min in 4%
paraformaldehyde-PBS. Coverslips were incubated
overnight in blocking solution with the following an-
tibodies: anti-Bassoon monoclonal mouse antibody
(ref. VAM-PS003, Stress Gen, USA) and rabbit
polyclonal sera against Synapsin (ref. 2312, Cell
Signaling, USA). Samples were incubated with a
fluorescence-conjugated secondary antibody in PBS
for 30 min. After that, coverslips were washed three
times in PBS and mounted using Mowiol (all sec-
ondary antibodies from Molecular Probes-Invitrogen,
USA). Stack images (pixel size 90 nm with 0.5 µm
Z step) were obtained with a Leica SP5 Confocal mi-
croscope (40x lens, 1. 3 NA). Percentage of synaptic
change is the average of different cultures under the
same experimental conditions. As a control, we used
sister untreated cultures growing in the same 24 well
multi plate.
A total of 13 individual images from three in-
dependent cultures has been analyzed. In Figure 6
we can observe that using a manual method to iden-
tify and count synapses, we obtain a mean of 24.12
synapses in control cultures and 16.74 in treated cul-
tures. The results obtained with SynapCountJ are
similar, there is a mean of 26.03 synapses in con-
trol cultures and 16.50 in the ones which have been
treated.
Notwithstanding the differences in the quantifica-
tion, in both procedures we obtain almost the same in-
hibition percentage, a 30.51% manually and 36.61%
automatically. This shows the suitability of Synap-
CountJ to count synapses, meaning a considerably re-
duction of the time employed in the manual process.
Namely, the manual analysis of an image takes ap-
proximately 5 minutes; of a batch, 1 hour; and, of
a complete study, 4 hours. Using SynapCountJ, the
time to analyze an image is 30 seconds; a batch, 2
minutes; and, a complete study, 6 minutes.
4 DISCUSSION
Up to the best of our knowledge, 4 tools have
been developed to quantify synapses and measure
synaptic density: Green and Red Puncta (Shi-
warski et al., 2014), Puncta Analyzer (Wark, 2013),
SynD (Schmitz et al., 2011) and SynPAnal (Daniel-
son and Lee, 2014) — a summary of the general fea-
tures of these tools can be seen in Table 1. The rest of
this section is devoted to compare SynapCountJ with
SynapCountJ: A Tool for Analyzing Synaptic Densities in Neurons
29

Table 1: General features of the analyzed software.
Software Language Underlying Tech-
nology
Types of Im-
ages
Technique for de-
tection
Green and Red Puncta Java ImageJ tiff Colocalization
Puncta Analyzer Java ImageJ2 tiff Colocalization
SynapCountJ Java ImageJ tiff and lif Colocalization
SynD Matlab Matlab tiff and lsm Brightness
SynPAnal Java tiff Brightness
Table 2: Features to quantify synapses and synaptic density of the analyzed software.
Software Detection of den-
drites
Threshold Batch
Processing
Dendrites
length
Density Export Save
Green and Red
Puncta
Not used X
Puncta Analyzer Manual ROI X X
SynapCountJ Manual X X X X X X
SynD Automatic X X X X X
SynPAnal Manual X X X X
these tools — such a comparison is summarized in
Table 2.
There are two approaches to locate synapses in an
RGB image either based on colocalization or bright-
ness. In the former, synapses are identified as the
colocalization of bright points in the red and green
channels — this is the approach followed by Green
and Red Puncta, Puncta Analyzer and SynapCountJ
— in the latter, synapses are the bright points of a re-
gion of an image — the approach employed in SynD
and SynPAnal. In both approaches, it is necessary a
threshold that can be manually adjusted to increase
(or decrease) the number of detected synapses; such a
functionality is supported by all the tools.
In the quantification of synapses from RGB im-
ages, it is instrumental to determine the region of in-
terest (i.e. the dendrites of the neurons where the
synapses are located); otherwise, the analysis will not
be precise due to noise coming from irrelevant regions
or the background of the image — this happens in
the Green and Red Puncta tool since it considers the
whole image for the analysis. Puncta Analyzer al-
lows the user to fix a rectangle containing the den-
drites of the neuron, but this is not completely pre-
cise since some regions of the rectangle might con-
tain points considered as synapses that do not belong
to the structure of the neuron. SynD is the only soft-
ware that automatically detects the dendrites of a neu-
ron; however, it can only be applied to neurons with
a cell-fill marker, and does not support the analysis
from specific regions, such as soma or distal den-
drites. SynapCountJ and SynPAnal provide the func-
tionality to manually draw the dendrites of the image;
allowing the user to designate the specific areas where
quantification is restricted.
The main output produced by all the available
tools is the number of synapses of a given image;
additionally, SynapCountJ, SynD and SynPAnal pro-
vides the length of the dendrites; and, SynapCountJ
is the only tool that outputs the synaptic density per
micron. All the tools but Green and Red Punctua can
export the results to an external file for storage and
further processing.
Finally, as we have explained in Subsection 2.2,
images obtained from the same biological experiment
usually have similar settings; hence, batch process-
ing might be useful. This functionality is featured by
SynapCountJ and SynD, and requires a previous step
of saving the configuration of an individual analysis.
SynPAnal does not support batch processing, but the
configuration of an individual analysis can be saved
to be later applied in other individual analysis.
As a summary, SynapCountJ is more complete
than the rest of available programs. It can use differ-
ent types of synaptic markers and can process batch
images. Furthermore, a differential feature of Synap-
CountJ is that it is based on a topological algorithm
(namely, computing the number of connected com-
ponents in a combinatorial structure), allowing us to
validate the correctness of our approach by means of
formal methods in software engineering.
5 CONCLUSIONS AND FURTHER
WORK
SynapCountJ is an ImageJ plugin that provides a
semi-automatic procedure to quantify synapses and
BIOIMAGING 2016 - 3rd International Conference on Bioimaging
30

measure synaptic density from immunofluorescence
images obtained from neuron cultures. This plugin
has been tested not only with neurons in development,
but also with the neuromuscular union of Drosophila;
therefore, it can be applied to the study of images that
contain two synaptic markers and a determined struc-
ture. The results obtained with SynapCountJ are con-
sistent with the results obtained manually; and Synap-
CountJ dramatically reduces the time required for the
quantification of synapses.
As further work, it remains the tasks of improv-
ing the usability of the plugin and including post-
processing tools to manually edit the obtained results.
Additionally, and since the final aim of our project
is the complete automation of the whole process, it
is necessary a procedure to automatically detect the
neuron morphology.
6 AVAILABILITY AND
SOFTWARE REQUIREMENTS
SynapCountJ is an ImageJ plugin that can be
downloaded, together with its documentation, from
http://imagejdocu.tudor.lu/doku.php?id=plugin:utiliti
es:synapsescountj:start. SynapCountJ is open source
and available for use under the GNU General Public
License. This plugin runs within both ImageJ and
Fiji (Schindelin et al., 2012) and has been tested on
Windows, Macintosh and Linux machines.
REFERENCES
Cuesto, G., Enriquez-Barreto, L., Caram
´
es, C., et al. (2011).
Phosphoinositide-3-kinase activation controls synap-
togenesis and spinogenesis in hippocampal neurons.
Journal of Neuroscience, 31(8):2721–2733.
Cuesto, G., Jord
´
an-
´
Alvarez, S., Enriquez-Barreto, L., et al.
(2015). GSK3β inhibition Promotes Synaptogenesis
in Drosophila and Mammalian Neurons. PlosOne,
10(3). doi=10.1371/journal.pone.0118475.
Danielson, E. and Lee, S. H. (2014). SynPAnal: Soft-
ware for Rapid Quantification of the Density and
Intensity of Protein Puncta from Fluorescence Mi-
croscopy Images of Neurons. PLoS ONE, 9(12).
doi=10.1371/journal.pone.0115298.
DaRocha-Souto, B., Scotton, T. C., Coma, M., et al. (2003).
Brain oligomeric β-amyloid but not total amyloid
plaque burden correlates with neuronal loss and astro-
cyte inflammatory response in amyloid precursor pro-
tein/tau transgenic mice. Journal of Neuropathology
& Experimental Neurology, 70(5):360–376.
Devices, M. (2015). Metamorph research imag-
ing. http://www.moleculardevices.com/systems/meta
morph-research-imaging.
Franco, B., Bogdanik, L., Bobinnec, Y., et al. (2004).
Shaggy, the homolog of glycogen synthase ki-
nase 3, controls neuromuscular junction growth in
Drosophila. Journal of Neuroscience, 24(29):6573–
6577.
Gonz
´
alez-D
´
ıaz, R. and Real, P. (2005). On the Cohomology
of 3D Digital Images. Discrete Applied Mathematics,
147(2–3):245–263.
Linkert, M., Rueden, C. T., Allan, C., et al. (2010). Meta-
data matters: access to image data in the real world.
The Journal of Cell Biology, 189(5):777–782.
Meijering, E., Jacob, M., Sarria, J. C. F., et al. (2004). De-
sign and Validation of a Tool for Neurite Tracing and
Analysis in Fluorescence Microscopy Images. Cytom-
etry Part A, 58(2):167–176.
Schindelin, J., Argand-Carreras, I., Frise, E., et al. (2012).
Fiji: an open-source platform for biological-image
analysis. Nature methods, 9(7):676–682.
Schmitz, S. K., Hjorth, J. J. J., Joemail, R. M. S., et al.
(2011). Automated analysis of neuronal morphology,
synapse number and synaptic recruitment. Journal of
Neuroscience Methods, 195(2):185–193.
Schneider, C., Rasband, W., and Eliceiri, K. (2012). NIH
Image to ImageJ. Nature Methods, 9:671–675.
S
´
egonne, F., Grimson, E., and Fischl, B. (2003). Topolog-
ical Correction of Subcortical Segmentation. In Pro-
ceedings of the 6th International conference on Med-
ical Image Computing and Computer Assisted Inter-
vention (MICCAI’03), volume 2879 of Lecture Notes
in Computer Science, pages 695–702.
Selkoe, D. J. (2002). Alzheimer’s diseases is a synaptic
failure. Science, 298(5594):789–791.
Shiwarski, D. J., Dagda, R. D., and Chu, C. T.
(2014). Green and red puncta colocalization.
http://imagejdocu.tudor.lu/doku.php?id=plugin:analy
sis:colocalization analysis macro for red and green
puncta:start.
Wark, B. (2013). Puncta analyzer v2.0. https://github.com/
physion/puncta-analyzer.
SynapCountJ: A Tool for Analyzing Synaptic Densities in Neurons
31
